This is the multi-page printable view of this section. Click here to print.
Documentation
- 1: Overview
- 2: Getting Started
- 2.1: First time users
- 2.2: Apache Installation
- 2.3: Docker Installation
- 2.4: Nginx Installation
- 2.5: Customize Installation
- 2.6: Upgrade OpenDataBio
- 3: API services
- 3.1: Quick reference
- 3.2: GET data
- 3.3: Post data
- 3.4: Put data
- 4: Concepts
- 4.1: Core Objects
- 4.2: Trait Objects
- 4.3: Data Access Objects
- 4.4: Auxiliary Objects
- 5: Contribution Guidelines
- 6: Tutorials
- 6.1: Getting data with OpenDataBio-R
- 6.2: Import data with R
- 6.2.1: Import Locations
- 6.2.2: Import BibReferences
- 6.2.3: Import Vernacular
- 6.2.4: Import Media
- 6.2.5: Import Taxons
- 6.2.6: Import Persons
- 6.2.7: Import Traits
- 6.2.8: Import Individuals & Vouchers
- 6.2.9: Import Measurements
1 - Overview
OpenDataBio is an opensource web-based platform designed to help researchers and organizations studying biodiversity in Tropical regions to collect, store, related and serve data. It is designed to accommodate many data types used in biological sciences and their relationships, particularly biodiversity and ecological studies, and serves as a data repository that allow users to download or request well-organized and documented research data.
Why?
Biodiversity studies frequently require the integration of a large amount of data, which require standardization for data use and sharing, and also continuous management and updates, particularly in Tropical regions where biodiversity is huge and poorly known.
OpenDataBio was designed based on the need to organize and integrate historical and current data collected in the Amazon region, taking into account field practices and data types used by ecologists and taxonomists.
OpenDataBio aim to facilitate the standardization and normalization of data, utilizing different API services available online, giving flexibility to user and user groups, and creating the necessary links among Locations, Taxons, Individuals, Vouchers and the Measurements and Media-files associated with them, while offering accessibility to the data through an API service, facilitating data distribution and analyses.
Main features
- Custom variables - the ability to define custom Traits, i.e. user defined variables of different types, including some special cases like Spectral Data, Colors, TaxonLinks and GeneBank. Measurements for such traits can be recorded for Individuals, Vouchers, Taxons and/or Locations.
- Taxons can be published or unpublished names (e.g. a morphotype), synonyms or valid names, and any node of the tree of life may be stored. Taxon insertion are checked against different nomenclature data sources (Tropicos, IPNI, MycoBank,ZOOBANK, GBIF), minimizing your search for correct spelling, authorship and synonyms.
- Locations are stored with their spatial Geometries, allowing location parent detection and spatial queries. Special location types, such as Plots and Transects can be defined, facilitating commonly used methods in biodiversity studies
- Data access control - data are organized in Datasets that permits to define an access policy (public, non-public) and a license for distribution of public datasets, becoming a self-contained dynamic data publication, versioned by the last edit date.
- Different research groups may use a single OpenDataBio installation, having total control over their particular research data edition and access, while sharing common libraries such as Taxonomy, Locations, Bibliographic References and Trait definitions.
- API to access data programatically - Tools for data exports and imports are provided through API services along with a API client in the R language, the OpenDataBio-R package.
- Autiting - the Activity Model audits changes in any record and downloads of full datasets, which are logged for history tracking.
- The BioCollection model allows administrators of Biological Collections to manage their Voucher records as well as user-requests, facilitating the interaction with users providing samples and data.
- A mobile data collector is planned with ODK or ODK-X
Learn more
- Getting Started: install OpenDataBio!
- Nginx Installation: standalone nginx setup.
- FAQs: Check out some example code!
2 - Getting Started
OpenDataBio is a web-based software supported in Debian, Ubuntu and Arch-Linux distributions of Linux and may be implemented in any Linux based machine. We have no plans for Windows support, but it may be easy to install on a Windows machine using Docker.
OpenDataBio is written in PHP and developed with the Laravel framework. It requires a web server (Apache or nginx), PHP and a SQL database – tested only with MySQL and MariaDB.
You may install OpenDataBio easily using the Docker files included in the distribution. The repository now includes a docker/prod profile and docker-compose.prod.yml so Docker can also be used for production deployments (with server-specific tuning and secrets management).
If you just want to test OpenDataBio locally, follow the Docker Installation.
Next steps
Prep for installation
- You may want to request a Tropicos.org API key for OpenDataBio to be able to retrieve taxonomic data from the Tropicos.org database. If not provided, mainly the GBIF nomenclatural service will be used;
- OpenDataBio sends emails to registered users, either to inform about a Job that has finished, to send data requests to dataset administrators or for password recovery. You may use a Google Email for this, but will need to change the account security options to allow OpenDataBio to use the account to send emails (you need to turn on the
Less secure app accessoption in the Gmail My Account Page and will need to create a cron job to keep this option alive). Therefore, create a dedicated email address for your installation. Check the “config/mail.php” file for more options on how to send e-mails.
2.1 - First time users
OpenDataBio is software to be used online. Local installations are for testing or development, although it could be used for a single-user production localhost environment.
User roles
If you have registered you need someone to assign you a full user role, so you can enter data.
- If you are installing, the first login to an OpenDataBio installation must be done with the default super-admin user:
admin@example.organdpassword1. These settings should be changed or the installation will be open to anyone reading the docs; - Self-registrations only grant access to datasets with privacy set to
registered usersand allows user do download data of open-access, but do not allow the user to edit nor add data; - Only full users can contribute with data.
- But only super admin can grant
full-user roleto registered users - different OpenDataBio installations may have different policies as to how you may gain full-user access. Here is not the place to find that info.
See also User Model.
Prep your full-user account
- Register yourself as Person and assign it as your user default person, creating a link between your user and yourself as collector.
- You need at least a dataset to enter your own data
- When becoming a full-user, a restricted-access Dataset and Project will be automatically created for you (your Workspaces). You may modify these entities to fit your personal needs.
- You may create as many Projects and Datasets as needed. So, understand how they work and which data they control access to.
Entering data
There three main ways to import data into OpenDataBio:
- One by one through the web Interface
- Using the OpenDataBio POST API services:
- importing from a spreadsheet file (CSV, XLXS or ODS) using the web Interface
- using the OpenDataBio R package client
- When using the OpenDataBio API services you must prep your data or file to import according to the field options of the POST verb for the specific ’endpoint’ your are trying to import.
See the Import data with R Tutorial for examples on how to import data with API.
Tips for entering data
- If first time entering data, you should use the web interface and create at least one record for each model needed to fit your needs. Then play with the privacy settings of your Workspace Dataset, and check whether you can access the data when logged in and when not logged in.
- Use Dataset for a self-contained set of data that should be distributed as a group. Datasets are dynamic publications, have author, data, and title.
- Although ODB attempt to minimize redundancy, giving users flexibility comes with a cost, and some definitions, like that of Traits or Persons may receive duplicated entries. So, care must be taken when creating such records. Administrators may create a ‘code of conduct’ for the users of an ODB installation to minimize such redundancy.
- Follow an order for importation of new data, starting from the libraries of common use. For example, you should first register Locations, Taxons, Persons, Traits and any other common library before importing Individuals or Measurements
- There is no need to import POINT locations before importing Individuals because ODB creates the location for you when you inform latitude and longitude, and will detect for you to which parent location your individual belongs to. However, if you want to validate your points (understand where such point location will placed), you may use the Location API with
querytypeparameter specified for this. - There are different ways to create PLOT and TRANSECT locations - see here Locations if that is your case
- Creating Taxons require only the specification of a
name- ODB will search nomenclature services for you, find the name, metadata and parents and import all of the them if needed. If you are importing published names, just inform this single attribute. Else, if the name is unpublished, you need to inform additional fields. So, separate the batch importation of published and unpublished names into two sets. - The
notesfield of any model is for both plain text or JSON object string formatted data. The Json option allows you to store custom structured data any model having thenotes field. You may, for example, store as notes some secondary fields from original sources when importing data, but may store any additional data that is not provided by the ODB database structure. Such data will not be validate by ODB and standardization of both tags and values depends on you. Json notes will be imported and exported as a JSON string, and will be presented in the interface as a formatted table; URLs in your Json will be presented as links.
2.2 - Apache Installation
These instructions are for an apache-based installation. For nginx, use Nginx Installation.
Server requirements
- The supported PHP version >= 8.2 (8.3 recommended)
- Web server: apache for this guide. For nginx, use Nginx Installation.
- It requires a SQL database, MySQL and MariaDB have been tested, but may also work with Postgres. Tested with MySQL 8.0 and MariaDB 10.6+.
- PHP extensions required:
openssl,pdo,pdo_mysql,mbstring,tokenizer,xml,dom,gd,exif,bcmath,zip,curl,redis. - Redis Server is required for queues and cache.
- Tectonic is used for LaTeX/PDF label generation.
- Pandoc is used to translate LaTeX code used in bibliographic references. It is not necessary for installation, but suggested for a better user experience.
- Requires Supervisor, which is needed background jobs
Create Dedicated User
The recommended way to install OpenDataBio for production is using a dedicated system user. In this instructions this user is odbserver.
Download OpenDataBio
Login as your Dedicated User and download or clone this software to where you want to install it.
Here we assume this is /home/odbserver/opendatabio so that the installation files will reside in this directory. If this is not your path, change below whenever it applies.
Download OpenDataBio
Prep the Server
First, install the prerequisite software: Apache, MySQL, PHP, Redis, Tectonic, Pandoc and Supervisor. On a Debian system, you need to install some PHP extensions as well and enable them:
sudo apt-get install software-properties-common
sudo add-apt-repository ppa:ondrej/php
sudo add-apt-repository ppa:ondrej/php ppa:ondrej/apache2
sudo add-apt-repository ppa:ondrej/php
sudo add-apt-repository ppa:ondrej/apache2
sudo apt-get install mysql-server redis-server tectonic php8.3 libapache2-mod-php8.3 php8.3-intl \
php8.3-mysql php8.3-sqlite3 php8.3-gd php8.3-cli pandoc \
php8.3-mbstring php8.3-xml php8.3-bcmath php8.3-zip php8.3-curl php8.3-redis \
supervisor
sudo a2enmod php8.3
sudo phpenmod mbstring
sudo phpenmod xml
sudo phpenmod dom
sudo phpenmod gd
sudo a2enmod rewrite
sudo a2ensite
sudo systemctl restart apache2.service
#To check if they are installed:
php -m | grep -E 'mbstring|cli|xml|gd|mysql|redis|bcmath|pcntl|zip'
tectonic --version
redis-server --version
Add the following to your Apache configuration.
- Change
/home/odbserver/opendatabioto your path (the files must be accessible by apache) - You may create a new file in the sites-available folder:
/etc/apache2/sites-available/opendatabio.confand place the following code in it.
touch /etc/apache2/sites-available/opendatabio.conf
echo '<IfModule alias_module>
Alias /opendatabio /home/odbserver/opendatabio/public/
Alias /fonts /home/odbserver/opendatabio/public/fonts
Alias /images /home/odbserver/opendatabio/public/images
Alias /build /home/odbserver/opendatabio/public/build
Alias /vendor/livewire /home/odbserver/opendatabio/public/vendor/livewire
<Directory "/home/odbserver/opendatabio/public">
Require all granted
AllowOverride All
</Directory>
</IfModule>' > /etc/apache2/sites-available/opendatabio.conf
This will cause Apache to redirect all requests for / to the correct folder, and also allow the provided .htaccess file to handle the rewrite rules, so that the URLs will be pretty. If you would like to access the file when pointing the browser to the server root, add the following directive as well:
RedirectMatch ^/$ /
Content Security Policy (CSP) for Apache
Configure CSP at the web server layer (not in Laravel files). Apply it first in report-only mode, inspect logs, then switch to enforced mode.
For nginx standalone installs, use Nginx Installation.
Apache: where to put it
- Enable required module:
sudo a2enmod headers
sudo systemctl restart apache2
- Edit your active vhost file (example):
sudo nano /etc/apache2/sites-available/opendatabio.conf
- Inside the correct
<VirtualHost ...>block (HTTP and/or HTTPS), add:
Header always set Content-Security-Policy-Report-Only "
default-src 'self';
base-uri 'self';
form-action 'self';
frame-ancestors 'self';
object-src 'none';
script-src 'self' 'unsafe-eval';
style-src 'self' 'unsafe-inline';
img-src 'self' data: https://server.arcgisonline.com https://*.tile.openstreetmap.org;
font-src 'self' data:;
connect-src 'self';
"
- Reload Apache:
sudo apachectl configtest
sudo systemctl reload apache2
Subpath installs (/opendatabio)
If your installation runs under a subpath (for example http://localhost/opendatabio), set in .env:
APP_URL=http://localhost/opendatabio
ASSET_URL=http://localhost/opendatabio
Then refresh generated assets and Livewire files:
php artisan livewire:publish --assets
php artisan optimize:clear
npm run build
Notes
https://server.arcgisonline.comandhttps://*.tile.openstreetmap.orgare needed for map tiles.unsafe-inline/unsafe-evalare temporary compatibility flags; remove after hardening templates/assets.- Keep
Report-Onlywhile tuning policy in production.
Configure your php.ini file. The installer may complain about missing PHP extensions, so remember to activate them in both the cli (/etc/php/8.3/cli/php.ini) and the web ini (/etc/php/8.3/fpm/php.ini) files for PHP!
Update the values for the following variables:
Find files:
php -i | grep 'Configuration File'
Change in them:
memory_limit should be at least 512M
post_max_size should be at least 30M
upload_max_filesize should be at least 30M
Something like:
[PHP]
allow_url_fopen=1
memory_limit = 512M
post_max_size = 100M
upload_max_filesize = 100M
Enable the Apache modules ‘mod_rewrite’ and ‘mod_alias’ and restart your Server:
sudo a2enmod rewrite
sudo a2ensite
sudo systemctl restart apache2.service
Mysql Charset and Collation
- You should add the following to your configuration file (mariadb.cnf or my.cnf), i.e. the Charset and Collation you choose for your installation must match that in the ‘config/database.php’
[mysqld]
character-set-client-handshake = FALSE #without this, there is no effect of the init_connect
collation-server = utf8mb4_unicode_ci
init-connect = "SET NAMES utf8mb4 COLLATE utf8mb4_unicode_ci"
character-set-server = utf8mb4
log-bin-trust-function-creators = 1
sort_buffer_size = 256M #large enough for geometry sort operations
[mariadb] or [mysql]
max_allowed_packet=100M
innodb_log_file_size=300M #no use for mysql
- If using MariaDB and you still have problems of type #1267 Illegal mix of collations, then check here on how to fix that,
Configure supervisord
Configure Supervisor, which is required for jobs. Create a file name opendatabio-worker.conf in the Supervisor configuration folder /etc/supervisor/conf.d/opendatabio-worker.conf with the following content:
touch /etc/supervisor/conf.d/opendatabio-worker.conf
echo ";--------------
[program:opendatabio-worker]
process_name=%(program_name)s_%(process_num)02d
command=php /home/odbserver/opendatabio/artisan queue:work --sleep=3 --tries=1 --timeout=0 --memory=512
autostart=true
autorestart=true
user=odbserver
numprocs=8
redirect_stderr=true
stdout_logfile=/home/odbserver/opendatabio/storage/logs/supervisor.log
;--------------" > /etc/supervisor/conf.d/opendatabio-worker.conf
Folder permissions
Folder and file permissions are important for securing a public server installation. If you don’t set them correctly, your site may be at risk.
- Folders
storageandbootstrap/cachemust be writable by the Server user (usually www-data). Set0755permission to these directories. - Config
.envfile requires0640permission. - This link has different ways to set up permissions for files and folders of a Laravel application. Below the preferred method:
cd /home/odbserver
#give write permissions to odbserver user and the apache user
sudo chown -R odbserver:www-data opendatabio
sudo find ./opendatabio -type f -exec chmod 644 {} \;
sudo find ./opendatabio -type d -exec chmod 755 {} \;
#in these folders the server stores data and files.
#Make sure their permission is correct
cd /home/odbserver/opendatabio
sudo chgrp -R www-data storage bootstrap/cache
sudo chmod -R ug+rwx storage bootstrap/cache
#make sure media folder has the correct permissions
sudo find ./storage/app/public/media -type f -exec chmod 664 {} \;
sudo find ./storage/app/public/media -type d -exec chmod 775 {} \;
#make sure the .env file has 640 permission
sudo chmod 640 ./.env
Install OpenDataBio
- Many Linux distributions (most notably Ubuntu and Debian) have different php.ini files for the command line interface and the Apache plugin. It is recommended to use the configuration file for Apache when running the install script, so it will be able to correctly point out missing extensions or configurations. To do so, find the correct path to the .ini file, and export it before using the
php installcommand.
For example,
export PHPRC=/etc/php/8.3/apache2/php.ini
The installation script will download the Composer dependency manager and all required PHP libraries listed in the
composer.jsonfile. However, if your server is behind a proxy, you should install and configure Composer independently. We have implemented PROXY configuration, but we are not using it anymore and have not tested properly (if you require adjustments, place an issue on GitLab).The script will prompt you configurations options, which are stored in the environment
.envfile in the application root folder.
You may, optionally, configure this file before running the installer:
- Create a
.envfile with the contents of the providedcp .env.example .env - Read the comments in this file and adjust accordingly.
- Make sure
assets_urlis correct for your deployment URL/subpath.
- Run the installer:
cd /home/odbserver/opendatabio
php install
- Build frontend assets after
.envis configured (required whenassets_urlis added or changed):
npm run build
- Seed data - the script above will ask if you want to install seed data for Locations and Taxons - seed data is version specific. Check the seed data repository version notes.
If the install script finishes with success, you’re good to go! Point your browser to http://localhost/opendatabio. The database migrations include an administrator account, with login admin@example.org and password password1. Change the password after installing.
Installation issues
There are countless possible ways to install the application, but they may involve more steps and configurations.
- if you browser return 500|SERVER ERROR you should look to the last error in
storage/logs/laravel.log. If you have ERROR: No application encryption key has been specified run:
php artisan key:generate
php artisan config:cache
- If you receive the error “failed to open stream: Connection timed out” while running the installer, this indicates a misconfiguration of your IPv6 routing. The easiest fix is to disable IPv6 routing on the server.
- If you receive errors during the random seeding of the database, you may attempt to remove the database entirely and rebuild it. Of course, do not run this on a production installation.
php artisan migrate:fresh
- You may also replace the Locations and Taxons tables with seed data after a fresh migration using:
php seedodb
Post-install configs
- If your import/export jobs are not being processed, make sure Supervisor is running
systemctl start supervisord && systemctl enable supervisord, and check the log files atstorage/logs/supervisor.log. - You can change several configuration variables for the application. The most important of those are probably set
by the installer, and include database configuration and proxy settings, but many more exist in the
.envandconfig/app.phpfiles. In particular, you may want to change the language, timezone and e-mail settings. Runphp artisan config:cacheafter updating the config files. - In order to stop search engine crawlers from indexing your database, add the following to your “robots.txt” in your server root folder (in Debian, /var/www/html):
User-agent: *
Disallow: /
Updating an existing Apache installation
Before updating, back up your database, .env, and storage/app/public/media.
Before running commands, review config diffs for the target version:
- Compare
.envwith.env.example(includingassets_url) - Check PHP settings (
php.iniin CLI and FPM/Apache) - Check Supervisor worker settings
- Put the application in maintenance mode:
cd /home/odbserver/opendatabio
php artisan down
- Update source code to the target version:
git fetch --tags
git checkout <target-tag-or-branch>
- Update dependencies and apply database migrations:
composer install --no-dev --optimize-autoloader
php artisan migrate:status
php artisan migrate --force
- Rebuild frontend assets after
.envupdates:
npm run build
- Refresh caches and restart queue workers:
php artisan optimize:clear
php artisan config:cache
php artisan queue:restart
echo "" > storage/logs/laravel.log
- Bring the application back online:
php artisan up
If the target version includes new environment variables (compare yours with the contents of .env.example), add them to .env before running asset/cache commands.
Storage & Backups
You may change storage configurations in config/filesystem.php, where you may define cloud based storage, which may be needed if have many users submitting media files, requiring lots of drive space.
- Data downloads are queue as jobs and a file is written in a temporary folder, and the file is deleted when the job is deleted by the user. This folder is defined as the
download diskin filesystem.php config file, which point tostorage/app/public/downloads. UserJobs web interface difficult navigation will force users to delete old jobs, but a cron cleaning job may be advisable to implement in your installation; - Media files are by default stored in the
media disk, which place files in folderstorage/app/public/media; - For regular configuration create both directories
storage/app/public/downloadsandstorage/app/public/mediawith writable permissions by the Server user, see below topic; - Remember to include media folder in a backup plan;
2.3 - Docker Installation
The easiest way to install and run OpenDataBio is using Docker and the docker configuration files provided, which contain the required configuration to run OpenDataBio. It uses nginx, MySQL, and Supervisor for queues
By default, the quick-start flow is optimized for development and local testing.
Production profile
OpenDataBio now ships a production-oriented Docker profile:
docker/prod/nginx.confdocker/prod/php.inidocker/prod/www.confdocker-compose.prod.yml
Run production compose with:
docker compose -f docker-compose.prod.yml build
docker compose -f docker-compose.prod.yml up -d
Key differences from dev:
- Uses
docker/prod/*nginx/php-fpm configs. - Removes source bind-mounts for app code.
- Disables phpMyAdmin by default (
dev-onlyprofile). - Publishes nginx on port
80(adjust if behind reverse proxy).
CSP in nginx (report-only) is included in docker/prod/nginx.conf. Keep report-only first, then enforce after validation.
Docker files
laravel-app/
----docker/*
----.env.docker
----docker-compose.yml
----Dockerfile
----Makefile
These are adapted from this link, where you find a production setting as well.
Installation
Download OpenDataBio
Prerequisites
- Docker with Compose plugin (
docker composev2). - Linux/mac: user in the docker group or run with
sudo. - Windows: Docker Desktop (WSL2/Hyper-V enabled).
Quick start (Linux/mac, requires make)
cd opendatabio
make docker-init # copies .env.docker, builds/starts, installs composer, key, migrates, storage:link
make docker-init SEED=1 # same as above + optional seed for Locations/Taxons
- After configuring
.env(or wheneverASSET_URLchanges), rebuild assets:
npm ci #may need this in production
npm run build
- App: http://localhost:8081 (user
admin@example.org/password1) - phpMyAdmin: http://localhost:8082
Windows (PowerShell)
cd opendatabio
powershell -ExecutionPolicy Bypass -File scripts/docker-init.ps1
# optional seed
powershell -ExecutionPolicy Bypass -File scripts/docker-init.ps1 -Seed
Manual commands (if you do not have make installed)
cp .env.docker .env
docker compose up -d
docker compose exec -T -u www-data laravel composer install --optimize-autoloader
docker compose exec -T -u www-data laravel php artisan key:generate --force
docker compose exec -T -u www-data laravel php artisan migrate --force
docker compose exec -T -u www-data laravel php artisan storage:link
Optional seed without make:
docker compose exec -T -u www-data laravel php getseeds
docker exec -i odb_mysql mysql -uroot -psecret odbdocker < storage/Location*.sql
docker exec -i odb_mysql mysql -uroot -psecret odbdocker < storage/Taxon*.sql
rm storage/Location*.sql storage/Taxon*.sql
Data persistence
The docker images may be deleted without loosing any data. The mysql tables are stored in a volume. You may change to a local path bind.
docker volume list
Using
The Makefile file contains the following commands to interact with the docker containers and odb.
Commands to build and create the app
make docker-init- copy.env.docker(if missing), build/start containers, install composer, key, migrate, storage:linkmake build- build containersmake key-generate- generate the app key and adds to .envmake composer-install- install php dependenciemake composer-update- update php dependenciesmake composer-dump-autoload- execute composer dump-autoload within containermake migrate- create or update the databasemake drop-migrate- delete and recreate the databasemake seed-odb- seed the database with locations and taxons
Commands to access the docker containers
make start- start all containersmake stop- stop all containersmake restart- restart all containersmake ssh- enter the main laravel app containermake ssh-mysql- enter the mysql container, so you may the log to the database usingmysql -uUSER -pPWDmake mysql- enter the docker mysql consolemake ssh-nginx- enter the nginx containermake ssh-supervisord- enter the supervisord container
Maintenance commands
make optimize- clean caches and log filesmake info- show app infomake logs- show laravel logsmake logs-mysql- show mysql logsmake logs-nginx- show nginx logsmake logs-supervisord- show supervisor logs
Deleting & rebuilding
If you have issues and changed the docker files, you may need to rebuild:
#delete all images without loosing data
make stop #first stop all
docker system prune -a #and accepts Yes
make build
make start
Updating an existing Docker installation
Before updating, back up your database and storage/app/public/media.
Before running commands, review config diffs for the target version:
- Compare
.envwith.env.example(includingassets_url) - Check PHP settings from the target profile (
docker/prod/php.inior your custom PHP config) - Check Supervisor settings (
docker/supervisord.confor your deployment equivalent)
- Update source code to the target version:
cd opendatabio
git fetch --tags
git checkout <target-tag-or-branch>
- Rebuild and restart containers:
make stop
make build
make start
- Update PHP dependencies and run database migrations:
make composer-install
make migrate
- Rebuild frontend assets after
.envupdates:
npm run build
- Refresh Laravel caches and restart queue workers:
make optimize
docker compose exec -T -u www-data laravel php artisan queue:restart
If the new version introduces changes in .env, add the new keys before asset/cache/worker refresh in production.
2.4 - Nginx Installation
These instructions are for an nginx-based installation. If you prefer Apache, use the Apache installation page.
Server requirements
- Supported PHP version >= 8.2 (8.3 recommended).
- Web server: nginx.
- SQL database: MySQL or MariaDB (tested with MySQL 8.0 and MariaDB 10.6+).
- Required PHP extensions:
openssl,pdo,pdo_mysql,mbstring,tokenizer,xml,dom,gd,exif,bcmath,zip,curl,redis. - Redis for queues/cache.
- Tectonic for label PDF generation.
- Pandoc for bibliographic rendering (recommended).
- Supervisor for background jobs.
Nginx site config
Create your site config file (example):
sudo nano /etc/nginx/sites-available/opendatabio
Use this base server block (adjust paths/domain):
server {
listen 80;
server_name your-domain.example;
root /home/odbserver/opendatabio/public;
index index.php index.html;
charset utf-8;
client_max_body_size 300M;
location / {
try_files $uri $uri/ /index.php?$query_string;
}
location ~ \.php$ {
try_files $uri =404;
fastcgi_split_path_info ^(.+\.php)(/.+)$;
fastcgi_pass unix:/var/run/php/php8.3-fpm.sock;
fastcgi_index index.php;
include fastcgi_params;
fastcgi_param SCRIPT_FILENAME $document_root$fastcgi_script_name;
fastcgi_param PATH_INFO $fastcgi_path_info;
fastcgi_read_timeout 300;
}
location ~ /\. {
deny all;
}
}
Enable and reload:
sudo ln -s /etc/nginx/sites-available/opendatabio /etc/nginx/sites-enabled/opendatabio
sudo nginx -t
sudo systemctl reload nginx
Content Security Policy (CSP)
Edit the same nginx site file and add inside the server { ... } block:
add_header Content-Security-Policy-Report-Only "
default-src 'self';
base-uri 'self';
form-action 'self';
frame-ancestors 'self';
object-src 'none';
script-src 'self' 'unsafe-eval';
style-src 'self' 'unsafe-inline';
img-src 'self' data: https://server.arcgisonline.com https://*.tile.openstreetmap.org;
font-src 'self' data:;
connect-src 'self';
" always;
Then reload:
sudo nginx -t
sudo systemctl reload nginx
Notes:
- Start with
Report-Only, then move to enforced CSP after validating logs. https://server.arcgisonline.comandhttps://*.tile.openstreetmap.orgare required for map tiles.
Subpath installs (/opendatabio)
If your installation runs under a subpath (for example http://localhost/opendatabio), set in .env:
APP_URL=http://localhost/opendatabio
ASSET_URL=http://localhost/opendatabio
Then refresh generated assets and Livewire files:
php artisan livewire:publish --assets
php artisan optimize:clear
npm run build
Shared application setup
To avoid repeating the same instructions, use these sections from Apache installation (they also apply to nginx deployments):
- PHP settings (php.ini) in Apache Installation
- Configure supervisord in Apache Installation
- Folder permissions in Apache Installation
- Install OpenDataBio in Apache Installation
- Post-install configs in Apache Installation
2.5 - Customize Installation
Simple changes that can be implemented in the layout of a OpenDataBio web site
Logo and BackGround Image
To replace the Navigation bar logo and the image of the landing page,
just put your image files replacing the files in /public/custom/ without changing their names.
Texts and Info
To change the welcome text of the landing page, change the values of the array keys in the following files:
/resources/lang/en/customs.php/resources/lang/pt-br/customs.php- Do not remove the entry keys. Set to null to suppress from appearing in the footer and landing page.
Local Documentation
You can add documentation in *.md format to the repository in files located in the following folders:
/resources/docs/en/*/resources/docs/pt/*
This space is reserved for administrators to set documentation and custom directives for users of a specific OpenDataBio installation. For example, this is a place to include a code of conduct for users, information on who to contact to become a full user, specific tutorials, and so on.
NavBar and Footer
- If you want to change the color of the top navigation bar and the footer,
just replace css Boostrap 5 class in the corresponding tags and files in folder
/resources/view/layout. - You may add additional html to the footer and navbar, change logo size, etc… as you wish.
2.6 - Upgrade OpenDataBio
Use this guide to upgrade an existing OpenDataBio installation with minimal downtime.
Before you start
- Read the target release notes and confirm any breaking changes.
- Back up at least:
- Database dump
.envstorage/app/public/media
- Compare current config files against target-version templates/settings:
.envagainst.env.example(includingassets_url)- Supervisor worker config (
/etc/supervisor/conf.d/opendatabio-worker.confor container equivalent) - PHP config (
php.inifor CLI and FPM/Apache)
- Plan a maintenance window for production.
Upgrade (Apache or nginx installation)
- Put application in maintenance mode:
cd /home/odbserver/opendatabio
php artisan down
- Update source code:
git fetch --tags
git checkout <target-tag-or-branch>
- Install dependencies and run migrations:
composer install --no-dev --optimize-autoloader
php artisan migrate:status
php artisan migrate --force
- Rebuild frontend assets after
.envchanges (required whenassets_urlchanges):
npm run build
- Refresh caches, clean logs, and restart workers:
php artisan optimize:clear
php artisan config:cache
php artisan queue:restart
systemctl restart supervisor.service, nginx+phpfpm or apache, mysql or mariadb
echo "" > storage/logs/laravel.log
echo "" > storage/logs/supervisor.log
- Bring app back online:
php artisan up
Upgrade (Docker installation)
- Update source code:
cd opendatabio
git fetch --tags
git checkout <target-tag-or-branch>
- Rebuild and restart containers:
make stop
make build
make start
- Install dependencies and run migrations:
make composer-install
make migrate
- Rebuild frontend assets after
.envchanges (required whenassets_urlchanges):
npm run build
- Refresh caches and restart workers:
make optimize
docker compose exec -T -u www-data laravel php artisan queue:restart
Environment variables
If the target version introduces new environment variables, compare .env with .env.example and add missing keys before asset/cache/worker commands in production.
Rollback strategy
If something fails after migration:
- Keep maintenance mode on.
- Restore database backup and
.env. - Checkout the previous known-good tag.
- Rebuild dependencies/containers and validate logs before
php artisan up.
3 - API services
Every OpenDataBio installation provide a API service, allowing users to GET data programmatically, and collaborators to POST new data into its database. The service is open access to public data, requires user authentication to POST data or GET data of restricted access.
The OpenDataBio API (Application Programming Interface -API) allows users to interact with an OpenDataBio database for exporting, importing and updating data without using the web-interface.
 The OpenDataBio R package is a client for this API, allowing the interaction with the data repository directly from R and illustrating the API capabilities so that other clients can be easily built.
The OpenDataBio R package is a client for this API, allowing the interaction with the data repository directly from R and illustrating the API capabilities so that other clients can be easily built.
The OpenDataBio API allows querying of the database, data importation and data edition (update) through a REST inspired interface. All API requests and responses are formatted in JSON.
The API call
A simple call to the OpenDataBio API has four independent pieces:
[HTTP-verb] + base-URL + endpoint + request-parameters
- HTTP-verb - either
GETfor exports orPOSTfor imports. - base-URL - the URL used to access your OpenDataBio server + plus
/api/v0. For, example,http://opendatabio.inpa.gov.br/api/v0 - endpoint - represents the object or collection of objects that you want to access, for example, for querying taxonomic names, the endpoint is “taxons”
- request-parameters - represent filtering and processing that should be done with the objects, and are represented in the API call after a question mark. For example, to retrieve only valid taxonomic names (non synonyms) end the request with
?valid=1.
The API call above can be entered in a browser to GET public access data. For example, to get the list of valid taxons from an OpenDataBio installation the API request could be:
https://opendb.inpa.gov.br/api/v0/taxons?valid=1&limit=10
When using the OpenDataBio R package this call would be odb_get_taxons(list(valid=1)).
A response would be something like:
{
"meta":
{
"odb_version":"0.9.1-alpha1",
"api_version":"v0",
"server":"http://opendb.inpa.gov.br",
"full_url":"https://opendb.inpa.gov.br/api/v0/taxons?valid=1&limit1&offset=100"},
"data":
[
{
"id":62,
"parent_id":25,
"author_id":null,
"scientificName":"Laurales",
"taxonRank":"Ordem",
"scientificNameAuthorship":null,
"namePublishedIn":"Juss. ex Bercht. & J. Presl. In: Prir. Rostlin: 235. (1820).",
"parentName":"Magnoliidae",
"family":null,
"taxonRemarks":null,
"taxonomicStatus":"accepted",
"ScientificNameID":"http:\/\/tropicos.org\/Name\/43000015 | https:\/\/www.gbif.org\/species\/407",
"basisOfRecord":"Taxon"
}]}
API Authentication
- Not required for getting any data with public access in the ODB database, which by default includes locations, taxons, bibliographic references, persons and traits.
- Authentication Required to GET any data that is not of public access, and is required to POST and PUT data.
- Authentication is done using an
API token, that can be found under your user profile on the web interface. The token is assigned to a single database user, and should not be shared, exposed, e-mailed or stored in version controls. - To authenticate against the OpenDataBio API, use the token in the “Authorization” header of the API request. When using the R client, pass the token to the
odb_configfunctioncfg = odb_config(token="your-token-here"). - The token controls the data you can get and can edit
Users will only have access to the data for which the user has permission and to any data with public access in the database, which by default includes locations, taxons, bibliographic references, persons and traits. Measurements, individuals, and Vouchers access depends on permissions understood by the users token.
API versions
The OpenDataBio API follows its own version number. This means that the client can expect to use the same code and get the same answers regardless of which OpenDataBio version that the server is running. All changes done within the same API version (>= 1) should be backward compatible. Our API versioning is handled by the URL, so to ask for a specific API version, use the version number between the base URL and endpoint:
https://opendatabio.inpa.gov.br/opendatabio/api/v1/taxons
https://opendatabio.inpa.gov.br/opendatabio/api/v2/taxons
3.1 - Quick reference
base-URL + ‘/api/v0/’ + endpoint + ‘?’ + request-parameters
https://opendb.inpa.gov.br/api/v0/taxons?valid=1&limit=2&offset=10
GET DATA (downloads)
Shared get-parameters
When multiple parameters are specified, they are combined with an AND operator. There is no OR parameter option in searches.
The limit and offset parameters can be used to divide your search into parts. Alternatively, use the save_job=T option and then download the data with the get_file=T parameter from the userjobs API.
Some parameters accept an asterisk as wildcard, so api/v0/taxons?name=Euterpe will return taxons with name exactly as “Euterpe”, while api/v0/taxons?name=Eut* will return names starting with “Eut”.
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
limit | No | Maximum number of records to return. | 100 |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
Endpoint parameters
| Endpoint | Description | Parameters |
|---|---|---|
| / | Tests your access/token. | — |
| bibreferences | Bibliographic references (GET lists, POST creates). | id, bibkey, biocollection, dataset, fields, job_id, limit, offset, save_job, search, taxon, taxon_root |
| biocollections | Biocollections (GET lists, POST creates). | id, acronym, fields, irn, job_id, limit, name, offset, save_job, search |
| datasets | Datasets and published dataset files (GET lists, POST creates via import job). | id, bibreference, fields, file_name, has_versions, include_url, limit, list_versions, name, offset, project, save_job, search, summarize, tag, tagged_with, taxon, taxon_root, traits |
| individuals | Individuals (GET lists, POST creates, PUT updates). | id, dataset, date_max, date_min, fields, job_id, limit, location, location_root, odbrequest_id, offset, person, project, save_job, tag, taxon, taxon_root, trait, vernacular |
| individual-locations | Occurrences for individuals with multiple locations (GET lists, POST/PUT upserts). | id, dataset, date_max, date_min, fields, individual, limit, location, location_root, offset, person, project, save_job, tag, taxon, taxon_root |
| languages | Lists available interface/data languages. | fields, limit, offset |
| locations | Locations (GET lists, POST creates, PUT updates). | id, adm_level, dataset, fields, job_id, lat, limit, location_root, long, name, offset, parent_id, project, querytype, root, save_job, search, taxon, taxon_root, trait |
| measurements | Trait measurements (GET lists, POST creates/imports via ImportMeasurements job, PUT bulk updates). | id, bibreference, dataset, date_max, date_min, fields, individual, job_id, limit, location, location_root, measured_id, measured_type, offset, person, project, save_job, taxon, taxon_root, trait, trait_type, voucher |
| media | Media metadata (GET lists, POST creates, PUT updates). | id, dataset, fields, individual, job_id, limit, location, location_root, media_id, media_uuid, offset, person, project, save_job, tag, taxon, taxon_root, uuid, voucher |
| persons | People (GET lists, POST creates, PUT updates). | id, abbrev, email, fields, job_id, limit, name, offset, save_job, search |
| projects | Projects (GET lists). | id, fields, job_id, limit, offset, save_job, search, tag |
| taxons | Taxonomic names (GET lists, POST creates). | id, bibreference, biocollection, dataset, external, fields, job_id, level, limit, location_root, name, offset, person, project, root, save_job, taxon_root, trait, valid, vernacular |
| traits | Trait definitions (GET lists, POST creates). | id, bibreference, categories, dataset, fields, job_id, language, limit, name, object_type, offset, save_job, search, tag, taxon, taxon_root, trait, type |
| vernaculars | Vernacular names (GET lists, POST creates). | id, fields, individual, job_id, limit, location, location_root, offset, save_job, taxon, taxon_root |
| vouchers | Voucher specimens (GET lists, POST creates, PUT updates). | id, bibreference, bibreference_id, biocollection, biocollection_id, collector, dataset, date_max, date_min, fields, individual, job_id, limit, location, location_root, main_collector, number, odbrequest_id, offset, person, project, save_job, taxon, taxon_root, trait, vernacular |
| userjobs | Background jobs (imports/exports) (GET lists). | id, fields, get_file, limit, offset, status |
| activities | Lists activity log entries. | id, description, fields, individual, language, limit, location, log_name, measurement, offset, save_job, subject, subject_id, taxon, taxon_root, voucher |
| tags | Tags/keywords (GET lists). | id, dataset, fields, job_id, language, limit, name, offset, project, save_job, search, trait |
POST DATA (imports)
Importing data from files through the web-interface require specifying the POST verb parameters of the ODB API
| Endpoint | Description | Parameters |
|---|---|---|
| bibreferences | Bibliographic references (GET lists, POST creates). | bibtex, doi |
| biocollections | Biocollections (GET lists, POST creates). | acronym, name |
| individuals | Individuals (GET lists, POST creates, PUT updates). | altitude, angle, biocollection, biocollection_number, biocollection_type, collector, dataset, date, distance, identification_based_on_biocollection, identification_based_on_biocollection_number, identification_date, identification_individual, identification_notes, identifier, latitude, location, location_date_time, location_notes, longitude, modifier, notes, tag, taxon, x, y |
| individual-locations | Occurrences for individuals with multiple locations (GET lists, POST/PUT upserts). | altitude, angle, distance, individual, latitude, location, location_date_time, location_notes, longitude, x, y |
| locations | Locations (GET lists, POST creates, PUT updates). | adm_level, altitude, azimuth, datum, geojson, geom, ismarine, lat, long, name, notes, parent, startx, starty, x, y |
| locations-validation | Validates coordinates against registered locations (POST). | latitude, longitude |
| measurements | Trait measurements (GET lists, POST creates/imports via ImportMeasurements job, PUT bulk updates). | bibreference, dataset, date, duplicated, link_id, location, notes, object_id, object_type, parent_measurement, person, trait_id, value |
| media | Media metadata (GET lists, POST creates, PUT updates). | collector, dataset, date, filename, latitude, license, location, longitude, notes, object_id, object_type, project, tags, title_en, title_pt |
| persons | People (GET lists, POST creates, PUT updates). | abbreviation, biocollection, email, full_name, institution |
| taxons | Taxonomic names (GET lists, POST creates). | author, author_id, bibreference, gbif, ipni, level, mobot, mycobank, name, parent, person, valid, zoobank |
| traits | Trait definitions (GET lists, POST creates). | bibreference, categories, description, export_name, link_type, name, objects, parent, range_max, range_min, tags, type, unit, value_length, wavenumber_max, wavenumber_min |
| vernaculars | Vernacular names (GET lists, POST creates). | citations, individuals, language, name, notes, parent, taxons, type |
| vouchers | Voucher specimens (GET lists, POST creates, PUT updates). | biocollection, biocollection_number, biocollection_type, collector, dataset, date, individual, notes, number |
| datasets | Datasets and published dataset files (GET lists, POST creates via import job). | description, license, name, privacy, project_id, title |
PUT DATA (updates)
Only the endpoints listed below can be updated using the API and only the listed PUT fields can be updated on each endpoint.
Field values are as explained for the POST API endpoints, except that in all cases you must also provide the id of the record to be updated.
| Endpoint | Description | Parameters |
|---|---|---|
| individuals | Individuals (GET lists, POST creates, PUT updates). | id, collector, dataset, date, identification_based_on_biocollection, identification_based_on_biocollection_number, identification_date, identification_individual, identification_notes, identifier, individual_id, modifier, notes, tag, taxon |
| individual-locations | Occurrences for individuals with multiple locations (GET lists, POST/PUT upserts). | id, altitude, angle, distance, individual, individual_location_id, latitude, location, location_date_time, location_notes, longitude, x, y |
| locations | Locations (GET lists, POST creates, PUT updates). | id, adm_level, altitude, datum, geom, ismarine, lat, location_id, long, name, notes, parent, startx, starty, x, y |
| measurements | Trait measurements (GET lists, POST creates/imports via ImportMeasurements job, PUT bulk updates). | id, bibreference, dataset, date, duplicated, link_id, location, measurement_id, notes, object_id, object_type, parent_measurement, person, trait_id, value |
| media | Media metadata (GET lists, POST creates, PUT updates). | id, collector, dataset, date, latitude, license, location, longitude, media_id, media_uuid, notes, project, tags, title_en, title_pt |
| persons | People (GET lists, POST creates, PUT updates). | id, abbreviation, biocollection, email, full_name, institution, person_id |
| vouchers | Voucher specimens (GET lists, POST creates, PUT updates). | id, biocollection, biocollection_number, biocollection_type, collector, dataset, date, individual, notes, number, voucher_id |
Nomenclature types
| Nomenclature types numeric codes | |
|---|---|
| NotType : 0 | Isosyntype : 8 |
| Type : 1 | Neotype : 9 |
| Holotype : 2 | Epitype : 10 |
| Isotype : 3 | Isoepitype : 11 |
| Paratype : 4 | Cultivartype : 12 |
| Lectotype : 5 | Clonotype : 13 |
| Isolectotype : 6 | Topotype : 14 |
| Syntype : 7 | Phototype : 15 |
Taxonomic ranks
| Code | Rank |
|---|---|
| -100 | clade |
| 0 | kingdom |
| 10 | subkingd. |
| 30 | div., phyl., phylum, division |
| 40 | subdiv. |
| 60 | cl., class |
| 70 | subcl., subclass |
| 80 | superord., superorder |
| 90 | ord., order |
| 100 | subord. |
| 120 | fam., family |
| 130 | subfam., subfamily |
| 150 | tr., tribe |
| 180 | gen., genus |
| 190 | subg., subgenus, sect. |
| 210 | section, sp., spec., species |
| 220 | subsp., subspecies |
| 240 | var., variety |
| 270 | f., fo., form |
3.2 - GET data
- The OpenDataBio-R package is a client for this API.
- Query examples in R are here;
- No authentication is needed to access data with a public access policy
- Authentication token is required only to get data with a non-public access policy
Shared GET parameters
When multiple parameters are specified, they are combined with an AND operator. There is no OR parameter option in searches.
The limit and offset parameters can be used to divide your search into parts. Alternatively, use the save_job=T option and then download the data with the get_file=T parameter from the userjobs API.
Some parameters accept an asterisk as wildcard, so api/v0/taxons?name=Euterpe will return taxons with name exactly as “Euterpe”, while api/v0/taxons?name=Eut* will return names starting with “Eut”.
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
limit | No | Maximum number of records to return. | 100 |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
GET endpoints
Quick links
- /
- bibreferences
- biocollections
- datasets
- individuals
- individual-locations
- languages
- locations
- measurements
- media
- persons
- projects
- taxons
- traits
- vernaculars
- vouchers
- userjobs
- activities
- tags
/ (GET)
Tests your access/token.
No parameters for this endpoint.
bibreferences (GET)
Bibliographic references (GET lists, POST creates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
bibkey | No | Bibreference key or list of keys. | ducke1953,mayr1992 |
biocollection | No | Biocollection id/name/acronym; returns references cited by vouchers in those collections. | INPA |
dataset | No | Dataset id or name; returns bibreferences linked to the dataset. | Forest1 |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
job_id | No | Job id to reuse affected ids or filter results from a job. | 1024 |
limit | No | Maximum number of records to return. | 100 |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
search | No | Full-text search on bibtex using boolean mode; spaces act as AND. | Amazon forest |
taxon | No | Taxon id or canonical name list; matches references linked to the taxon. | Ocotea guianensis or 120,455 |
taxon_root | No | Taxon id/name including descendants. | Lauraceae |
Fields returned
Fields (simple): id, bibkey, year, author, title, doi, url, bibtex
Fields (all): id, bibkey, year, author, title, doi, url, bibtex
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 2,
"bibkey": "Riberiroetal1999FloraDucke",
"year": 1999,
"author": "José Eduardo Lahoz Da Silva Ribeiro and Michael John Gilbert Hopkins and Alberto Vicentini and Cynthia Anne Sothers and Maria Auxiliadora Da Silva Costa and Joneide Mouzinho De Brito and Maria Anália Duarte De Souza and Lúcia Helena Pinheiro Martins and Lúcia Garcez Lohmann and Paulo Apóstolo Costa Lima Assunção and Everaldo Da Costa Pereira and Cosme Fernandes Da Silva and Mariana Rabello Mesquita and Lilian Costa Procópio",
"title": "Flora Da Reserva Ducke: Guia De Identificação Das Plantas Vasculares De Uma Floresta De Terra Firme Na Amazônica Central",
"doi": null,
"url": null,
"bibtex": "@Article{Riberiroetal1999FloraDucke,\r\n title = {Flora da Reserva Ducke: Guia de Identifica{\\c{c}}{\\~a}o das Plantas Vasculares de uma Floresta de Terra Firme na Amaz{\\^o}nica Central},\r\n author = {José Eduardo Lahoz da Silva Ribeiro and Michael John Gilbert Hopkins and Alberto Vicentini and Cynthia Anne Sothers and Maria Auxiliadora da Silva Costa and Joneide Mouzinho de Brito and Maria Anália Duarte de Souza and Lúcia Helena Pinheiro Martins and Lúcia Garcez Lohmann and Paulo Apóstolo Costa Lima Assunç{ã}o and Everaldo da Costa Pereira and Cosme Fernandes da Silva and Mariana Rabello Mesquita and Lilian Costa Procópio},\r\n journal = {Flora da Reserva Ducke: Guia de Identifica{\\c{c}}{\\~a}o das Plantas Vasculares de uma Floresta de Terra Firme na Amaz{\\^o}nica Central},\r\n year = {1999},\r\n publisher = {INPA-DFID Manaus},\r\n pages = {819p},\r\n}"
},
{
"id": 3,
"bibkey": "Sutter2006female",
"year": 2006,
"author": "D. Merino Sutter and P. I. Forster and P. K. Endress",
"title": "Female Flowers And Systematic Position Of Picrodendraceae (Euphorbiaceae S.l., Malpighiales)",
"doi": "10.1007/s00606-006-0414-0",
"url": "http://dx.doi.org/10.1007/s00606-006-0414-0",
"bibtex": "@article{Sutter2006female,\n author = {D. Merino Sutter and P. I. Forster and P. K. Endress},\n year = {2006},\n title = {Female flowers and systematic position of Picrodendraceae (Euphorbiaceae s.l., Malpighiales)},\n issn = {0378-2697 | 1615-6110},\n issue = {1-4},\n url = {http://dx.doi.org/10.1007/s00606-006-0414-0},\n doi = {10.1007/s00606-006-0414-0},\n volume = {261},\n page = {187-215},\n journal = {Plant Systematics and Evolution},\n journal_short = {Plant Syst. Evol.},\n published = {Springer Science and Business Media LLC}\n}"
}
]
}biocollections (GET)
Biocollections (GET lists, POST creates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
acronym | No | Biocollection acronym. | INPA |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
irn | No | Index Herbariorum IRN for filtering biocollections. | 123456 |
job_id | No | Job id to reuse affected ids or filter results from a job. | 1024 |
limit | No | Maximum number of records to return. | 100 |
name | No | Exact biocollection name (string). | Instituto Nacional de Pesquisas da Amazônia |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
search | No | Full-text search parameter. | Silva |
Fields returned
Fields (simple): id, acronym, name, irn
Fields (all): id, acronym, name, irn, country, city, address
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 1,
"acronym": "INPA",
"name": "Instituto Nacional de Pesquisas da Amazônia",
"irn": 124921,
"country": null,
"city": null,
"address": null
},
{
"id": 2,
"acronym": "SPB",
"name": "Universidade de São Paulo",
"irn": 126324,
"country": null,
"city": null,
"address": null
}
]
}datasets (GET)
Datasets and published dataset files (GET lists, POST creates via import job).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
bibreference | No | Bibreference id or bibkey. | 34 or ducke1953 |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
file_name | No | Dataset version file name to download. | 2_Organisms.csv |
has_versions | No | When 1, returns only datasets that have public versions. | 1 |
include_url | No | When 1 with list_versions, include file download URL. | 1 |
limit | No | Maximum number of records to return. | 100 |
list_versions | No | If true, lists dataset version files for given id(s). | 1 |
name | No | Translatable trait name. Accepts a plain string or a JSON map of language codes to names. | {"en":"Height","pt-br":"Altura"} |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
project | No | Project id or acronym. | PDBFF or 2 |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
search | No | Full-text search parameter. | Silva |
summarize | No | Dataset id to return content/taxonomic/trait summaries. | 3 |
tag | No | Individual tag/number/code. | A-1234 |
tagged_with | No | Tag ids (comma) or text to filter datasets by tags (supports id list or full-text). | 12,13 or canopy leaf |
taxon | No | Taxon id or canonical full name list. | Licaria cannela or 456,789 |
taxon_root | No | Taxon id/name including descendants. | Lauraceae |
traits | No | Trait ids list (comma-separated) for filtering datasets. | 12,15 |
Fields returned
Fields (simple): id, name, title, projectName, description, notes, contactEmail, taggedWidth, uuid
Fields (all): id, name, title, projectName, notes, privacyLevel, policy, description, measurements_count, contactEmail, taggedWidth, uuid
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 4,
"name": "PDBFF-FITO 1ha core plots 1-10cm dbh - TREELETS",
"title": "Arvoretas (1cm>DAP",
"projectName": "Projeto Dinâmica Biológica de Fragmentos Florestais (PDBFF-Data)",
"notes": null,
"privacyLevel": "Restrito a usuários autorizados",
"policy": null,
"description": "Contém o único censo de árvores de pequeno porte 1-10cm de diâmetro nas parcelas de 1ha do PDBFF, em 11 das 69 de parcelas permanentes de 1ha do Programa de Monitoramento de Plantas do PDBFF.",
"measurements_count": null,
"contactEmail": "example",
"taggedWidth": "Parcelas florestais | PDBFF | Fitodemográfico",
"uuid": "e1d8ce8d-4847-11f0-8e9f-9cb654b86224"
}
]
}individuals (GET)
Individuals (GET lists, POST creates, PUT updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
dataset | No | Dataset id/name, filter records that belong to the dataset informed | 3 or FOREST1 |
date_max | No | Inclusive end date (YYYY-MM-DD) compared against individual date. | 2024-12-31 |
date_min | No | Inclusive start date (YYYY-MM-DD) compared against individual date. | 2020-01-01 |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
job_id | No | Job id to reuse affected ids or filter results from a job. | 1024 |
limit | No | Maximum number of records to return. | 100 |
location | No | Location id/name list; matches individuals at those exact locations. | Parcela 25ha or 55,60 |
location_root | No | Location id/name; includes descendants of the informed locations. | Parcela 25ha get subplots in this case |
odbrequest_id | No | Request id to filter individuals linked to that ODB request. | 12 |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
person | No | Collector person id/name/email list; filters main/associated collectors. | Silva, J.B. or 23,10 |
project | No | Project id/name; matches records whose dataset belongs to the project. | PDBFF |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
tag | No | Individual tag/number filter; supports list separated by comma. | A-123,2001 |
taxon | No | Taxon id/name list; matches identification taxon only (no descendants). | Licaria guianensis,Minquartia guianensis or 456,457 |
taxon_root | No | Taxon id/name list; includes descendants of each taxon. | Lauraceae,Fabaceae or 10,20 |
trait | No | Trait id list; only used together with dataset to filter by measurements. | 12,15 |
vernacular | No | Vernacular id/name list to match linked vernaculars. | castanha|12 |
Fields returned
Fields (simple): id, basisOfRecord, organismID, recordedByMain, recordNumber, recordedDate, family, scientificName, identificationQualifier, identifiedBy, dateIdentified, locationName, locationParentName, decimalLatitude, decimalLongitude, x, y, gx, gy, angle, distance, datasetName
Fields (all): id, basisOfRecord, organismID, recordedByMain, recordNumber, recordedDate, recordedBy, scientificName, scientificNameAuthorship, taxonPublishedStatus, genus, family, identificationQualifier, identifiedBy, dateIdentified, identificationRemarks, identificationBiocollection, identificationBiocollectionReference, locationName, higherGeography, decimalLatitude, decimalLongitude, georeferenceRemarks, locationParentName, x, y, gx, gy, angle, distance, organismRemarks, datasetName, uuid
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 306246,
"basisOfRecord": "Organism",
"organismID": "2639_Spruce_1852",
"recordedByMain": "Spruce, R.",
"recordNumber": "2639",
"recordedDate": "1852-10",
"recordedBy": "Spruce, R.",
"scientificName": "Ecclinusa lanceolata",
"scientificNameAuthorship": "(Mart. & Eichler) Pierre",
"taxonPublishedStatus": "published",
"genus": "Ecclinusa",
"family": "Sapotaceae",
"identificationQualifier": "",
"identifiedBy": "Spruce, R.",
"dateIdentified": "1852-10-00",
"identificationRemarks": "",
"identificationBiocollection": null,
"identificationBiocollectionReference": null,
"locationName": "São Gabriel da Cachoeira",
"higherGeography": "São Gabriel da Cachoeira < Amazonas < Brasil",
"decimalLatitude": 1.1841927,
"decimalLongitude": -66.80167715,
"georeferenceRemarks": "decimal coordinates are the CENTROID of the footprintWKT geometry",
"locationParentName": "Amazonas",
"x": null,
"y": null,
"gx": null,
"gy": null,
"angle": null,
"distance": null,
"organismRemarks": "prope Panure ad Rio Vaupes Amazonas, Brazil",
"datasetName": "Exsicatas LABOTAM",
"uuid": "c01000f0-f437-11ef-b90b-9cb654b86224"
}
]
}individual-locations (GET)
Occurrences for individuals with multiple locations (GET lists, POST/PUT upserts).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
dataset | No | Dataset id/name; filters by dataset of the linked individual. | FOREST1 |
date_max | No | Upper bound date/time; compares date_time or individual date when empty. | 2024-12-31 |
date_min | No | Lower bound date/time; compares date_time or individual date when empty. | 2020-01-01 |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
individual | No | Individual id list whose occurrences will be returned. | 12,44 |
limit | No | Maximum number of records to return. | 100 |
location | No | Location id or name. | Parcela 25ha or 55 |
location_root | No | Location id/name with descendants included. | Amazonas or 10 |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
person | No | Collector person id/name/email list; filters by individual collectors. | J.Silva|23 |
project | No | Project id/name; matches occurrences whose individual belongs to datasets in project. | PDBFF |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
tag | No | Individual tag/number list; matches by individuals.tag column | A-123,B-2 |
taxon | No | Taxon id or canonical full name list. | Licaria cannela or 456,789 |
taxon_root | No | Taxon id/name including descendants. | Lauraceae |
Fields returned
Fields (simple): id, individual_id, basisOfRecord, occurrenceID, organismID, recordedDate, locationName, higherGeography, decimalLatitude, decimalLongitude, x, y, angle, distance, minimumElevation, occurrenceRemarks, scientificName, family, datasetName
Fields (all): id, individual_id, basisOfRecord, occurrenceID, organismID, scientificName, family, recordedDate, locationName, higherGeography, decimalLatitude, decimalLongitude, georeferenceRemarks, x, y, angle, distance, minimumElevation, occurrenceRemarks, organismRemarks, datasetName
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 306244,
"individual_id": 306246,
"basisOfRecord": "Occurrence",
"occurrenceID": "2639_Spruce_1852.1852-10",
"organismID": "2639_Spruce_1852",
"scientificName": "Ecclinusa lanceolata",
"family": "Sapotaceae",
"recordedDate": "1852-10",
"locationName": "São Gabriel da Cachoeira",
"higherGeography": "Brasil > Amazonas > São Gabriel da Cachoeira",
"decimalLatitude": 1.1841927,
"decimalLongitude": -66.80167715,
"georeferenceRemarks": "decimal coordinates are the CENTROID of the footprintWKT geometry",
"x": null,
"y": null,
"angle": null,
"distance": null,
"minimumElevation": null,
"occurrenceRemarks": null,
"organismRemarks": "prope Panure ad Rio Vaupes Amazonas, Brazil",
"datasetName": "Exsicatas LABOTAM"
}
]
}languages (GET)
Lists available interface/data languages.
| Parameter | Required | Description | Example |
|---|---|---|---|
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
limit | No | Maximum number of records to return. | 100 |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 1,
"code": "en",
"name": "English",
"is_locale": 1,
"created_at": null,
"updated_at": null
}
]
}locations (GET)
Locations (GET lists, POST creates, PUT updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
adm_level | No | One or more adm_level codes | 10,100 |
dataset | No | Dataset id/name; expands to all locations used by that dataset. | FOREST1 |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
job_id | No | Job id to reuse affected ids or filter results from a job. | 1024 |
lat | No | Latitude (decimal degrees) used with querytype. | -3.11 |
limit | No | Maximum number of records to return. | 100 |
location_root | No | Alias of root for compatibility. | Amazonas |
long | No | Longitude (decimal degrees) used with querytype. | -60.02 |
name | No | Exact name match; accepts list of names or ids. | Manaus or 10 |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
parent_id | No | Parent id for hierarchical queries. | 210 |
project | No | Project id or acronym. | PDBFF or 2 |
querytype | No | When lat/long are provided: exact|parent|closest geometric search. | parent |
root | No | Location id/name; returns it and all descendants and related locations | Amazonas or "Parque Nacional do Jaú" ... |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
search | No | Prefix search on name (SQL LIKE name%). | Mana search for names that starts "mana" |
taxon | No | Taxon id/name list; filters locations by linked identifications. | Euterpe precatoria |
taxon_root | No | Taxon id/name list; includes descendants when filtering linked identifications. | Euterpe - finds alls records that belongs to this genus |
trait | No | Trait id/name; only works together with dataset to filter by measurements. | DBH |
Fields returned
Fields (simple): id, basisOfRecord, locationName, adm_level, country_adm_level, x, y, startx, starty, distance_to_search, parent_id, parentName, higherGeography, footprintWKT, locationRemarks, decimalLatitude, decimalLongitude, georeferenceRemarks, geodeticDatum
Fields (all): id, basisOfRecord, locationName, adm_level, country_adm_level, x, y, startx, starty, distance_to_search, parent_id, parentName, higherGeography, footprintWKT, locationRemarks, decimalLatitude, decimalLongitude, georeferenceRemarks, geodeticDatum
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 27297,
"basisOfRecord": "Location",
"locationName": "Parcela 1105",
"adm_level": 100,
"country_adm_level": "Parcela",
"x": "100.00",
"y": "100.00",
"startx": null,
"starty": null,
"distance_to_search": null,
"parent_id": 27277,
"parentName": "Fazenda Esteio",
"higherGeography": "Brasil > Amazonas > Rio Preto da Eva > Fazenda Esteio > Parcela 1105",
"footprintWKT": "POLYGON((-59.81371985 -2.42215752,-59.81360263 -2.42126619,-59.81270751 -2.42136656,-59.81282469 -2.42225788,-59.81371985 -2.42215752))",
"locationRemarks": "source: Polígono desenhado a partir das coordenadas de GPS dos vértices; georeferencedBy: Diogo Martins Rosa & Ana Andrade; fundedBy: Edital CNPq-Brasil/LBA 458027/2013-8; geometryBy: Alberto Vicentini; geometryDate: 2021-09-29; warning: Conflito com polígono da UC de 2021. Este polígono deveria ter a mesma geometria do polígono correspondente que faz parte da UC ARIE PDBFF, mas como ele foi gerado pelas coordenadas de campo, foi mantida essa geometria. A UC, portanto, não protege adequadamente essa parcela de monitoramento.",
"decimalLatitude": -2.42215752,
"decimalLongitude": -59.81371985,
"georeferenceRemarks": "decimal coordinates are the START POINT in footprintWKT geometry",
"geodeticDatum": null
}
]
}measurements (GET)
Trait measurements (GET lists, POST creates/imports via ImportMeasurements job, PUT bulk updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
bibreference | No | Bibreference id or bibkey. | 34 or ducke1953 |
dataset | No | Dataset id or acronym. | 3 or FOREST1 |
date_max | No | Filter records occurring on/before this date (YYYY-MM-DD). | 2024-12-31 |
date_min | No | Filter records occurring on/after this date (YYYY-MM-DD). | 2020-01-01 |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
individual | No | Individual id, uuid or organismID (fullname). | 4521 or 2ff0e884-3d33 |
job_id | No | Job id to reuse affected ids or filter results from a job. | 1024 |
limit | No | Maximum number of records to return. | 100 |
location | No | Location id or name. | Parcela 25ha or 55 |
location_root | No | Location id/name with descendants included. | Amazonas or 10 |
measured_id | No | Measurement filter: id of the measured object (matches measured_type). | 4521 |
measured_type | No | Measurement filter: class name of measured object (Individual, Location, Taxon, Voucher, Media). | Media |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
person | No | Person id, abbreviation, full name or email (supports lists with | or ;). | J.Silva|M.Costa |
project | No | Project id or acronym. | PDBFF or 2 |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
taxon | No | Taxon id or canonical full name list. | Licaria cannela or 456,789 |
taxon_root | No | Taxon id/name including descendants. | Lauraceae |
trait | No | Trait id or export_name filter. | DBH |
trait_type | No | Filter measurements by trait type code. | 1 |
voucher | No | Voucher id for filtering measurements. | 102 |
Fields returned
Fields (simple): id, basisOfRecord, measured_type, measured_id, measurementType, measurementValue, measurementUnit, measurementDeterminedBy, measurementDeterminedDate, scientificName, datasetName, family, sourceCitation
Fields (all): id, basisOfRecord, measured_type, measured_id, measurementType, measurementValue, measurementUnit, measurementDeterminedDate, measurementDeterminedBy, measurementRemarks, resourceRelationship, resourceRelationshipID, relationshipOfResource, scientificName, family, datasetName, measurementMethod, sourceCitation, measurementLocationId, measurementParentId, decimalLatitude, decimalLongitude
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 1,
"basisOfRecord": "MeasurementsOrFact",
"measured_type": "App\\Models\\Individual",
"measured_id": 86947,
"measurementType": "treeDbh",
"measurementValue": 13,
"measurementUnit": "cm",
"measurementDeterminedDate": "1979-11-14",
"measurementDeterminedBy": "Menezes, J.F. | Bahia, R.P. | Lima, J. | Santos, R.M. | Ferreira, A.J.C. | Cardoso, Romeu M.",
"measurementRemarks": null,
"resourceRelationship": null,
"resourceRelationshipID": "1202-1371_Menezes_1979",
"relationshipOfResource": "measurement of",
"scientificName": "Paramachaerium ormosioides",
"family": "Fabaceae",
"datasetName": "Censos 01 - PDBFF-FITO ForestPlots - 1979-1980",
"measurementMethod": "Name: Diameter at breast height - DBH | Definition:Diameter at breast height,, i.e. ca. 1.3 meters from the base of the trunk",
"sourceCitation": null,
"measurementLocationId": 28280,
"measurementParentId": null,
"decimalLatitude": -2.40371599,
"decimalLongitude": -59.87090972
}
]
}media (GET)
Media metadata (GET lists, POST creates, PUT updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
dataset | No | Dataset id or acronym. | 3 or FOREST1 |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
individual | No | Individual id, uuid or organismID (fullname). | 4521 or 2ff0e884-3d33 |
job_id | No | Job id to reuse affected ids or filter results from a job. | 1024 |
limit | No | Maximum number of records to return. | 100 |
location | No | Location id or name. | Parcela 25ha or 55 |
location_root | No | Location id/name with descendants included. | Amazonas or 10 |
media_id | No | Media numeric id. | 88 |
media_uuid | No | Media UUID. | a3f0a4ac-6b5b-11ed-b8c0-0242ac120002 |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
person | No | Person id, abbreviation, full name or email (supports lists with | or ;). | J.Silva|M.Costa |
project | No | Project id or acronym. | PDBFF or 2 |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
tag | No | Individual tag/number/code. | A-1234 |
taxon | No | Taxon id or canonical full name list. | Licaria cannela or 456,789 |
taxon_root | No | Taxon id/name including descendants. | Lauraceae |
uuid | No | — | |
voucher | No | Voucher id for filtering measurements. | 102 |
Fields returned
Fields (simple): id, model_type, model_id, basisOfRecord, recordedBy, recordedDate, dwcType, resourceRelationship, resourceRelationshipID, relationshipOfResource, scientificName, family, datasetName, projectName, taggedWith, accessRights, license, file_name, file_url, citation, uuid
Fields (all): id, model_type, model_id, basisOfRecord, recordedBy, recordedDate, dwcType, resourceRelationship, resourceRelationshipID, relationshipOfResource, scientificName, family, datasetName, projectName, taggedWith, accessRights, bibliographicCitation, license, file_name, file_url, citation, uuid, bibtex, userName, created_at
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 20211,
"model_type": "App\\Models\\Individual",
"model_id": 111785,
"basisOfRecord": "MachineObservation",
"recordedBy": "Francisco Javier Farroñay Pacaya",
"recordedDate": "2025-03-09",
"dwcType": "StillImage",
"resourceRelationship": "Organism",
"resourceRelationshipID": "3402-1134_Pereira_1986",
"relationshipOfResource": "StillImage of ",
"scientificName": "Sacoglottis guianensis",
"family": "Humiriaceae",
"datasetName": "Unknown dataset",
"projectName": "Projeto Dinâmica Biológica de Fragmentos Florestais",
"taggedWith": "Folha abaxial",
"accessRights": "Open access.",
"bibliographicCitation": "Sacoglottis guianensis (Humiriaceae). (2025). By Francisco Javier Farroñay Pacaya. Collection: Pereira, M.J.R. #3402-1134 on 1986-01-24, from Quadrante 52, Parcela 3402-3, Reserva 3402, Cabo Frio, Fazenda Porto Alegre, Amazonas, Brasil (PDBFF). Project: PDBFF-Data. Instituto Nacional de Pesquisas da Amazônia (INPA), Manaus, Amazonas, Brasil. Type: Image. License: CC-BY-NC-SA 4.0. uuid: inpa-odb-3f139ba4-f22b-42d8-9e74-c340309061c2, url: http://localhost/opendatabio",
"license": "CC-BY-NC-SA 4.0",
"file_name": "67ce28cd76f4a.jpg",
"file_url": "http://localhost/opendatabio/storage/media/20211/67ce28cd76f4a.jpg",
"citation": "Sacoglottis guianensis (Humiriaceae). (2025). By Francisco Javier Farroñay Pacaya. Collection: Pereira, M.J.R. #3402-1134 on 1986-01-24, from Quadrante 52, Parcela 3402-3, Reserva 3402, Cabo Frio, Fazenda Porto Alegre, Amazonas, Brasil (PDBFF). Project: PDBFF-Data. Instituto Nacional de Pesquisas da Amazônia (INPA), Manaus, Amazonas, Brasil. Type: Image. License: CC-BY-NC-SA 4.0. uuid: inpa-odb-3f139ba4-f22b-42d8-9e74-c340309061c2, url: http://localhost/opendatabio",
"uuid": "3f139ba4-f22b-42d8-9e74-c340309061c2",
"bibtex": "@misc{Farronay_2025_20211,\n{\n \"title\": \" Sacoglottis guianensis (Humiriaceae)\",\n \"year\": \"(2025)\",\n \"author\": \"Francisco Javier Farroñay Pacaya\",\n \"howpublished\": \"{http:\\/\\/localhost\\/opendatabio\\/media\\/uuid\\/3f139ba4-f22b-42d8-9e74-c340309061c2}\",\n \"license\": \"CC-BY-NC-SA 4.0\",\n \"note\": \"Type: Image; Collection: Pereira, M.J.R. #3402-1134 on 1986-01-24, from Quadrante 52, Parcela 3402-3, Reserva 3402, Cabo Frio, Fazenda Porto Alegre, Amazonas, Brasil (PDBFF); Coordinates: POINT(-59.91500315727877 -2.3929141688648765); License: CC-BY-NC-SA 4.0; Project: PDBFF-Data.; Accessed: 2026-02-04\",\n \"publisher\": \"Instituto Nacional de Pesquisas da Amazônia (INPA), Manaus, Amazonas, Brasil\"\n}\n}",
"userName": "example",
"created_at": "2025-03-09T23:48:29.000000Z"
}
]
}persons (GET)
People (GET lists, POST creates, PUT updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
abbrev | No | Abbreviation search for persons. | Silva, J.B, Pilco, M.V. |
email | No | Email address. | user@example.org |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
job_id | No | Job id to reuse affected ids or filter results from a job. | 1024 |
limit | No | Maximum number of records to return. | 100 |
name | No | Translatable trait name. Accepts a plain string or a JSON map of language codes to names. | {"en":"Height","pt-br":"Altura"} |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
search | No | Full-text search parameter. | Silva |
Fields returned
Fields (simple): id, full_name, abbreviation, emailAddress, institution, notes
Fields (all): id, full_name, abbreviation, emailAddress, institution, notes
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 3127,
"full_name": "Raimundo Afeganistão",
"abbreviation": "AFEGANISTÃO, R.",
"emailAddress": null,
"institution": null,
"notes": "PDBFF"
},
{
"id": 14,
"full_name": "Maria de Fátima Agra",
"abbreviation": "Agra, M.F.",
"emailAddress": null,
"institution": null,
"notes": null
},
{
"id": 15,
"full_name": "J. L. A. Aguiar Jr",
"abbreviation": "Aguiar Jr., J.L.A.",
"emailAddress": null,
"institution": null,
"notes": null
}
]
}projects (GET)
Projects (GET lists).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
job_id | No | Job id to reuse affected ids or filter results from a job. | 1024 |
limit | No | Maximum number of records to return. | 100 |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
search | No | Full-text search parameter. | Silva |
tag | No | Individual tag/number/code. | A-1234 |
Fields returned
Fields (simple): id, acronym, name, description
Fields (all): id, acronym, name, description, pages, urls, created_at, updated_at
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 1,
"acronym": "PDBFF-Data",
"name": "Projeto Dinâmica Biológica de Fragmentos Florestais",
"description": "Este espaço agrega conjuntos de dados de monitoramentos e pesquisas realizadas nas áreas amostrais do PDBFF, localizadas na Área de Relevante Interesse Ecológico - ARIE PDBFF.",
"pages": {
"en": null,
"pt-br": null
},
"urls": [
{
"url": "https://alfa-pdbff.site/",
"label": null,
"icon": "fa-solid fa-globe"
}
],
"created_at": "2022-10-31T07:01:18.000000Z",
"updated_at": "2023-11-17T21:08:55.000000Z"
}
]
}taxons (GET)
Taxonomic names (GET lists, POST creates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
bibreference | No | Bibreference id or bibkey. | 34 or ducke1953 |
biocollection | No | Biocollection id, name or acronym. | INPA |
dataset | No | Dataset id or acronym. | 3 or FOREST1 |
external | No | Flag to include external ids (Tropicos, IPNI, etc.). | 1 |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
job_id | No | Job id to reuse affected ids or filter results from a job. | 1024 |
level | No | Taxon rank code or string. | 210 or species |
limit | No | Maximum number of records to return. | 100 |
location_root | No | Location id/name with descendants included. | Amazonas or 10 |
name | No | Translatable trait name. Accepts a plain string or a JSON map of language codes to names. | {"en":"Height","pt-br":"Altura"} |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
person | No | Person id, abbreviation, full name or email (supports lists with | or ;). | J.Silva|M.Costa |
project | No | Project id or acronym. | PDBFF or 2 |
root | No | Root id for hierarchical queries (taxon or location). | 120 |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
taxon_root | No | Taxon id/name including descendants. | Lauraceae |
trait | No | Trait id or export_name filter. | DBH |
valid | No | When 1, return only valid taxon names. | 1 |
vernacular | No | Vernacular id or name list used to filter individuals. | castanha|12 |
Fields returned
Fields (simple): id, parent_id, author_id, scientificName, taxonRank, scientificNameAuthorship, namePublishedIn, parentName, family, taxonRemarks, taxonomicStatus, scientificNameID, basisOfRecord
Fields (all): id, senior_id, parent_id, author_id, scientificName, taxonRank, scientificNameAuthorship, namePublishedIn, parentName, family, higherClassification, taxonRemarks, taxonomicStatus, acceptedNameUsage, acceptedNameUsageID, parentNameUsage, scientificNameID, basisOfRecord
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 16332,
"senior_id": null,
"parent_id": 16331,
"author_id": null,
"scientificName": "Aiouea grandifolia",
"taxonRank": "Species",
"scientificNameAuthorship": "van der Werff",
"namePublishedIn": null,
"parentName": "Aiouea",
"family": "Lauraceae",
"higherClassification": "Eukaryota > Plantae > Viridiplantae > Embryophytes > Spermatopsida > Angiosperms > Magnoliidae > Laurales > Lauraceae > Aiouea",
"taxonRemarks": null,
"taxonomicStatus": "accepted",
"acceptedNameUsage": null,
"acceptedNameUsageID": null,
"parentNameUsage": "Aiouea",
"scientificNameID": "https://tropicos.org/Name/17806050 | https://www.gbif.org/species/4175896",
"basisOfRecord": "Taxon"
}
]
}traits (GET)
Trait definitions (GET lists, POST creates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
bibreference | No | Bibreference id or bibkey. | 34 or ducke1953 |
categories | No | Trait categories JSON list with lang/rank/name/description. | [{\"lang\":\"en\",\"rank\":1,\"name\":\"small\"}] |
dataset | No | Dataset id or acronym. | 3 or FOREST1 |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
job_id | No | Job id to reuse affected ids or filter results from a job. | 1024 |
language | No | Language id/code/name | en or 1 or english or portuguese |
limit | No | Maximum number of records to return. | 100 |
name | No | Translatable trait name. Accepts a plain string or a JSON map of language codes to names. | {"en":"Height","pt-br":"Altura"} |
object_type | No | Measured object type: Individual, Location, Taxon, Voucher, or Media. | Individual |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
search | No | Full-text search parameter. | Silva |
tag | No | Individual tag/number/code. | A-1234 |
taxon | No | Taxon id or canonical full name list. | Licaria cannela or 456,789 |
taxon_root | No | Taxon id/name including descendants. | Lauraceae |
trait | No | Trait id or export_name filter. | DBH |
type | No | Generic type parameter (trait type code or vernacular type such as use/generic/etimology). | use or 10 |
Fields returned
Fields (simple): id, type, typename, export_name, unit, range_min, range_max, link_type, value_length, name, description, objects, measurementType, categories
Fields (all): id, type, typename, export_name, measurementType, measurementUnit, range_min, range_max, link_type, value_length, name, description, objects, measurementMethod, MeasurementTypeBibkeys, TaggedWith, categories
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 206,
"type": 1,
"typename": "QUANT_REAL",
"export_name": "treeDbh",
"measurementType": "treeDbh",
"measurementUnit": "cm",
"range_min": 0.1,
"range_max": 700,
"link_type": null,
"value_length": null,
"name": "Diâmetro à altura do peito – DAP",
"description": "Diâmetro à altura do peito, i.e. medido a ca. 1.3m desde a base do caule",
"objects": "App\\Models\\Individual | App\\Models\\Voucher | App\\Models\\Location | App\\Models\\Taxon | App\\Models\\Media",
"measurementMethod": "Name: Diameter at breast height - DBH | Definition:Diameter at breast height,, i.e. ca. 1.3 meters from the base of the trunk",
"MeasurementTypeBibkeys": "",
"TaggedWith": "",
"categories": null
},
{
"id": 207,
"type": 1,
"typename": "QUANT_REAL",
"export_name": "treeDbhPom",
"measurementType": "treeDbhPom",
"measurementUnit": "m",
"range_min": 0,
"range_max": 15,
"link_type": null,
"value_length": null,
"name": "Ponto de medição do DAP",
"description": "Ponto de medição do DAP, necessário quando impossível medir a 1.3 m",
"objects": "App\\Models\\Individual",
"measurementMethod": "Name: DBH Point of Measurement | Definition:DAP measuring height, necessary when impossible to measure at 1.3 m",
"MeasurementTypeBibkeys": "",
"TaggedWith": "",
"categories": null
},
{
"id": 524,
"type": 2,
"typename": "CATEGORICAL",
"export_name": "stemType",
"measurementType": "stemType",
"measurementUnit": null,
"range_min": null,
"range_max": null,
"link_type": null,
"value_length": null,
"name": "Tipo de fuste",
"description": "Tipo de fuste",
"objects": "App\\Models\\Voucher | App\\Models\\Individual | App\\Models\\Taxon",
"measurementMethod": "Name: Type of stem | Definition:Type of stem | Categories: CategoryName: Main stem | Definition:The main trunk, usually the thickest. | CategoryName: Secondary stem | Definition:A secondary trunk, there is a thicker one, which defines the area better. A shoot below 1.3 m high is a secondary trunk.",
"MeasurementTypeBibkeys": "",
"TaggedWith": "",
"categories": [
{
"id": 12990,
"name": "Fuste principal",
"description": "O tronco principal, geralmente o mais grosso.",
"rank": 1,
"belongs_to_trait": "stemType"
},
{
"id": 12991,
"name": "Fuste secundário",
"description": "Um tronco secundário, há outro mais grosso, que define melhor a área. Um rebroto abaixo de 1.3 m de altura é um tronco secundário.",
"rank": 2,
"belongs_to_trait": "stemType"
}
]
}
]
}vernaculars (GET)
Vernacular names (GET lists, POST creates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
individual | No | Individual id, uuid or organismID (fullname). | 4521 or 2ff0e884-3d33 |
job_id | No | Job id to reuse affected ids or filter results from a job. | 1024 |
limit | No | Maximum number of records to return. | 100 |
location | No | Location id or name. | Parcela 25ha or 55 |
location_root | No | Location id/name with descendants included. | Amazonas or 10 |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
taxon | No | Taxon id or canonical full name list. | Licaria cannela or 456,789 |
taxon_root | No | Taxon id/name including descendants. | Lauraceae |
Fields returned
Fields (simple): id, name, languageName, notes, locationsList, taxonsList, individualsList, citationsArray
Fields (all): id, name, languageName, languageCode, notes, taxonsList, taxonsListArray, individualsList, individualsListArray, locationsList, locationsListArray, variantsList, variantsListArray, citationsArray, createdBy, created_at, updated_at
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 1,
"name": "itaúba-preta",
"languageName": "Portuguese",
"languageCode": "pt-br",
"notes": null,
"taxonsList": "Mezilaurus duckei",
"taxonsListArray": [
{
"id": 19774,
"scientificName": "Mezilaurus duckei",
"family": "Lauraceae"
}
],
"individualsList": "7739_Macedo_2023",
"individualsListArray": [
{
"id": 510747,
"uuid": "c75d9233-f437-11ef-b90b-9cb654b86224",
"organismId": "7739_Macedo_2023",
"scientificName": "Mezilaurus duckei",
"family": "Lauraceae"
}
],
"locationsList": null,
"locationsListArray": [],
"variantsList": "itaúba",
"variantsListArray": [
{
"id": 2,
"name": "itaúba",
"languageName": "Tupi",
"languageCode": "tup"
}
],
"citationsArray": [
{
"id": 2,
"citation": "ita = pedra; uba = árvore; preta em referência a cor da madeira",
"bibreference_id": null,
"bibreference_name": null,
"notes": null,
"type": "etimology",
"createdBy": "example"
}
],
"createdBy": "example",
"created_at": "2025-12-15T20:17:16.000000Z",
"updated_at": "2025-12-15T21:04:53.000000Z"
}
]
}vouchers (GET)
Voucher specimens (GET lists, POST creates, PUT updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
bibreference | No | Bibreference id or bibkey. | 34 or ducke1953 |
bibreference_id | No | BibReference id list for voucher filtering. | 10,11 |
biocollection | No | Biocollection id, name or acronym. | INPA |
biocollection_id | No | Biocollection id list for voucher filtering. | 1,5 |
collector | No | Collector(s) id, abbreviation, name or email. Use | or ; to separate multiple people; first is main collector. | J.Silva|M.Costa |
dataset | No | Dataset id or acronym. | 3 or FOREST1 |
date_max | No | Filter records occurring on/before this date (YYYY-MM-DD). | 2024-12-31 |
date_min | No | Filter records occurring on/after this date (YYYY-MM-DD). | 2020-01-01 |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
individual | No | Individual id, uuid or organismID (fullname). | 4521 or 2ff0e884-3d33 |
job_id | No | Job id to reuse affected ids or filter results from a job. | 1024 |
limit | No | Maximum number of records to return. | 100 |
location | No | Location id or name. | Parcela 25ha or 55 |
location_root | No | Location id/name with descendants included. | Amazonas or 10 |
main_collector | No | Boolean (1) to filter vouchers by main collector only. | 1 |
number | No | Collector number/code (voucher/individual tag when different from individual). | 1234A |
odbrequest_id | No | Filter individuals linked to a given request id. | 12 |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
person | No | Person id, abbreviation, full name or email (supports lists with | or ;). | J.Silva|M.Costa |
project | No | Project id or acronym. | PDBFF or 2 |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
taxon | No | Taxon id or canonical full name list. | Licaria cannela or 456,789 |
taxon_root | No | Taxon id/name including descendants. | Lauraceae |
trait | No | Trait id or export_name filter. | DBH |
vernacular | No | Vernacular id or name list used to filter individuals. | castanha|12 |
Fields returned
Fields (simple): id, individual_id, basisOfRecord, occurrenceID, organismID, collectionCode, catalogNumber, typeStatus, recordedByMain, recordNumber, recordedDate, recordedBy, scientificName, family, identificationQualifier, identifiedBy, dateIdentified, identificationRemarks, locationName, decimalLatitude, decimalLongitude, occurrenceRemarks, datasetName
Fields (all): id, individual_id, basisOfRecord, occurrenceID, organismID, collectionCode, catalogNumber, typeStatus, recordedByMain, recordNumber, recordedDate, recordedBy, scientificName, scientificNameAuthorship, taxonPublishedStatus, genus, family, identificationQualifier, identifiedBy, dateIdentified, identificationRemarks, locationName, higherGeography, decimalLatitude, decimalLongitude, georeferenceRemarks, occurrenceRemarks, datasetName, uuid
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 72209,
"individual_id": 306246,
"basisOfRecord": "PreservedSpecimens",
"occurrenceID": "2639.Spruce.K.K000640463",
"organismID": "2639_Spruce_1852",
"collectionCode": "K",
"catalogNumber": "K000640463",
"typeStatus": "Tipo",
"recordedByMain": "Spruce, R.",
"recordNumber": "2639",
"recordedDate": "1852-10",
"recordedBy": "Spruce, R.",
"scientificName": "Ecclinusa lanceolata",
"scientificNameAuthorship": "(Mart. & Eichler) Pierre",
"taxonPublishedStatus": "published",
"genus": "Ecclinusa",
"family": "Sapotaceae",
"identificationQualifier": "",
"identifiedBy": "Spruce, R.",
"dateIdentified": "1852-10-00",
"identificationRemarks": "",
"locationName": "São Gabriel da Cachoeira",
"higherGeography": "Brasil > Amazonas > São Gabriel da Cachoeira",
"decimalLatitude": 1.1841927,
"decimalLongitude": -66.80167715,
"georeferenceRemarks": "decimal coordinates are the CENTROID of the footprintWKT geometry",
"occurrenceRemarks": "OrganismRemarks = prope Panure ad Rio Vaupes Amazonas, Brazil",
"datasetName": "Exsicatas LABOTAM",
"uuid": "6302316f-2b48-43b5-816b-005df70d15c9"
}
]
}userjobs (GET)
Background jobs (imports/exports) (GET lists).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
get_file | No | When 1 and used with userjobs id, returns job prepared file. | 1 |
limit | No | Maximum number of records to return. | 100 |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
status | No | Job status filter (Submitted, Processing, Success, Failed, Cancelled). | Success |
Fields returned
Fields (simple): id, dispatcher, status, percentage, created_at, affected_ids, affected_model
Fields (all): id, dispatcher, status, percentage, created_at, updated_at, affected_ids, affected_model, log
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 23652,
"dispatcher": "App\\Jobs\\BatchUpdateIndividuals",
"status": "Success",
"percentage": "100%",
"created_at": "2025-12-10T14:36:25.000000Z",
"updated_at": "2025-12-10T14:36:28.000000Z",
"affected_ids": [
86136,
57362,
85053,
72256,
74543
],
"affected_model": "App\\Models\\Individual",
"log": "[]"
}
]
}activities (GET)
Lists activity log entries.
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
description | No | Translatable description text. Accepts a plain string or a JSON map of language codes to descriptions. | {"en":"Tree height at breast height","pt-br":"Altura da árvore à altura do peito"} |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
individual | No | Individual id, uuid or organismID (fullname). | 4521 or 2ff0e884-3d33 |
language | No | Language id/code/name | en or 1 or english or portuguese |
limit | No | Maximum number of records to return. | 100 |
location | No | Location id or name. | Parcela 25ha or 55 |
log_name | No | Activity log name filter. | default |
measurement | No | Activity filter: measurement id. | 55 |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
subject | No | Activity filter: subject type (class basename). | Individual |
subject_id | No | Activity filter: subject id. | 12 |
taxon | No | Taxon id or canonical full name list. | Licaria cannela or 456,789 |
taxon_root | No | Taxon id/name including descendants. | Lauraceae |
voucher | No | Voucher id for filtering measurements. | 102 |
Fields returned
Fields (simple): id, log_name, description, subject_type, subject_name, subject_id, modified_by, properties, created_at, updated_at
Fields (all): id, log_name, description, subject_type, subject_id, subject_name, modified_by, properties, created_at, updated_at
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"field_key": "taxon_id",
"field": "Taxon",
"old_value": "Burseraceae",
"new_value": "Protium hebetatum forma.b.fito",
"id": 1411696,
"log_name": "individual",
"description": "identification updated",
"subject_type": "App\\Models\\Individual",
"subject_id": 301705,
"subject_name": null,
"modified_by": "example"
},
{
"field_key": "person_id",
"field": "Person",
"old_value": "Macedo, M.T.S",
"new_value": "Pilco, M.V.",
"id": 1411696,
"log_name": "individual",
"description": "identification updated",
"subject_type": "App\\Models\\Individual",
"subject_id": 301705,
"subject_name": null,
"modified_by": "example"
},
{
"field_key": "notes",
"field": "Notes",
"old_value": "Identificação feita em campo, anotada na planilha de dados.",
"new_value": null,
"id": 1411696,
"log_name": "individual",
"description": "identification updated",
"subject_type": "App\\Models\\Individual",
"subject_id": 301705,
"subject_name": null,
"modified_by": "example"
},
{
"field_key": "date",
"field": "Date",
"old_value": "2022-06-17",
"new_value": "2022-11-23",
"id": 1411696,
"log_name": "individual",
"description": "identification updated",
"subject_type": "App\\Models\\Individual",
"subject_id": 301705,
"subject_name": null,
"modified_by": "example"
}
]
}tags (GET)
Tags/keywords (GET lists).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. | 1,2,3 |
dataset | No | Dataset id or acronym. | 3 or FOREST1 |
fields | No | Comma separated list of the fields to include in the response or special words all/simple/raw, default to simple | id,scientificName or all |
job_id | No | Job id to reuse affected ids or filter results from a job. | 1024 |
language | No | Language id/code/name | en or 1 or english or portuguese |
limit | No | Maximum number of records to return. | 100 |
name | No | Translatable trait name. Accepts a plain string or a JSON map of language codes to names. | {"en":"Height","pt-br":"Altura"} |
offset | No | The starting position of the record set to be exported. Used together with limit to limit results. | 10000 |
project | No | Project id or acronym. | PDBFF or 2 |
save_job | No | If 1, save the results as file to download later via userjobs + get_file = T | 1 |
search | No | Full-text search parameter. | Silva |
trait | No | Trait id or export_name filter. | DBH |
Fields returned
Fields (simple): id, name, description
Fields (all): id, name, description, counts
Response example
{
"meta": {
"odb_version": "0.10.0-alpha1",
"api_version": "v0",
"server": "http://localhost/opendatabio"
},
"data": [
{
"id": 11,
"name": "Folhas adaxial",
"description": "Images of the adaxial surface of leaves",
"counts": {
"Media": 1852,
"Project": 0,
"Dataset": 0,
"ODBTrait": 0
}
},
{
"id": 12,
"name": "Folha forma",
"description": "Imagem mostrando uma folha ou o formato da folha.",
"counts": {
"Media": 713,
"Project": 0,
"Dataset": 0,
"ODBTrait": 0
}
},
{
"id": 13,
"name": "Frutos",
"description": "Imagens com frutos",
"counts": {
"Media": 2595,
"Project": 0,
"Dataset": 0,
"ODBTrait": 0
}
}
]
}3.3 - Post data
Importing data
- The OpenDataBio-R package is a client for this API and lets you import, download and update data from R objects.
- Examples of importing from R are here;
- Data can be imported via the web interface using spreadsheets, where the column names are the POST parameters listed on this page
- An authentication token is required for all POST requests
Structured custom data in the notes field
The notes field of any model is for plain text or a text formatted as a JSON object containing structured data. Json allows you to store custom structured data in any model that has the notes field. For example, you may want to store secondary fields from source datasets during import, or any additional data not provided by the OpenDataBio database structure. This data is not validated by OpenDataBio and the standardization of tags and values is up to you. Json notes will be imported and exported as JSON text and will be presented in the interface as a formatted table; URLs in your Json will be presented as links in this table.
POST endpoints
Quick links
- bibreferences
- biocollections
- individuals
- individual-locations
- locations
- locations-validation
- measurements
- media
- persons
- taxons
- traits
- vernaculars
- vouchers
- datasets
bibreferences (POST)
Bibliographic references (GET lists, POST creates).
| Parameter | Required | Description | Example |
|---|---|---|---|
bibtex | No | BibTeX formatted reference string. (Provide doi or bibtex.) | @article{mykey,...} |
doi | No | DOI number or URL. (Provide doi or bibtex.) | 10.1234/abcd.2020.1 |
biocollections (POST)
Biocollections (GET lists, POST creates).
| Parameter | Required | Description | Example |
|---|---|---|---|
acronym | Yes | Biocollection acronym. | INPA |
name | Yes | Translatable trait name. Accepts a plain string or a JSON map of language codes to names. | {"en":"Height","pt-br":"Altura"} |
individuals (POST)
Individuals (GET lists, POST creates, PUT updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
altitude | No | Elevation in meters. | 75 |
angle | No | Azimuth from reference point, in degrees. | 45 |
biocollection | No | Biocollection id, name or acronym. | INPA |
biocollection_number | No | Catalogue number/code inside the biocollection. | 12345 |
biocollection_type | No | Nomenclatural type code or name. | Holotype or 2 |
collector | Yes | Collector(s) id, abbreviation, name or email. Use | or ; to separate multiple people; first is main collector. | J.Silva|M.Costa |
dataset | Yes | Dataset id or acronym. | 3 or FOREST1 |
date | Yes | Date (YYYY-MM-DD) or incomplete date (e.g. 1888-05-NA) or array with year/month/day. (At least the year must be provided.) | 2024-05-20 or {\"year\":1888,\"month\":5} |
distance | No | Distance from reference point in meters. | 12.5 |
identification_based_on_biocollection | No | Biocollection name/id used as reference for identification. | INPA |
identification_based_on_biocollection_number | No | CatalogNumber in the reference biocollection | 8765 |
identification_date | No | Identification date (full or incomplete). | 2023-06-NA |
identification_individual | No | ID/organismID of another individual that provides the taxonomic identification for the individual’s record | 3245 or REC-123 |
identification_notes | No | Notes for the identification. | Checked with microscope |
identifier | No | Person(s) responsible for identification; accept id, abbreviation, name or email; separate multiple with | or ;. | Costa, A.|Lima, B. or 1|2|3 or Adolpho Ducke|José Ramos|Paulo Apóstolo Costa Lima Assunção |
latitude | No | Latitude in decimal degrees (negative for south). (Required when location is not provided.) | -3.101 |
location | No | Location id or name. (Required when latitude/longitude are not provided.) | Parcela 25ha or 55 |
location_date_time | No | Date or date+time for the occurrence/location event. (Required when adding multiple locations or when different from individual date.) | 2023-08-14 12:30:00 |
location_notes | No | Notes for the occurrence/location entry. | Near trail marker 10 |
longitude | No | Longitude in decimal degrees (negative for west). (Required when location is not provided.) | -60.12 |
modifier | No | Identification modifier code/name (s.s.=1, s.l.=2, cf.=3, aff.=4, vel aff.=5). | 3 |
notes | No | Free text or JSON notes field. | {\"expedition\":\"2024-01\",\"tag\":\"P1\"} |
tag | Yes | Individual tag/number/code. | A-1234 |
taxon | No | Taxon id or canonical full name list. | Licaria cannela or 456,789 |
x | No | X coordinate for plots or individual position. | 12.3 |
y | No | Y coordinate for plots or individual position. | 8.7 |
individual-locations (POST)
Occurrences for individuals with multiple locations (GET lists, POST/PUT upserts).
| Parameter | Required | Description | Example |
|---|---|---|---|
altitude | No | Elevation in meters. | 75 |
angle | No | Azimuth from reference point, in degrees. | 45 |
distance | No | Distance from reference point in meters. | 12.5 |
individual | Yes | Individual id, uuid or organismID (fullname). | 4521 or 2ff0e884-3d33 |
latitude | No | Latitude in decimal degrees (negative for south). (Required when location is not provided.) | -3.101 |
location | No | Location id or name. (Required when latitude/longitude are not provided.) | Parcela 25ha or 55 |
location_date_time | Yes | Date or date+time for the occurrence/location event. | 2023-08-14 12:30:00 |
location_notes | No | Notes for the occurrence/location entry. | Near trail marker 10 |
longitude | No | Longitude in decimal degrees (negative for west). (Required when location is not provided.) | -60.12 |
x | No | X coordinate for plots or individual position. | 12.3 |
y | No | Y coordinate for plots or individual position. | 8.7 |
locations (POST)
Locations (GET lists, POST creates, PUT updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
adm_level | Yes | Location administrative level code (e.g. 100=plot, 10=country). | 100 |
altitude | No | Elevation in meters. | 75 |
azimuth | No | Azimuth (degrees) used to build plot/transect geometry when location is a POINT. | 90 |
datum | No | Spatial datum/projection string. | EPSG:4326-WGS 84 |
geojson | No | Single GeoJSON Feature with geometry and at least name + adm_level properties. | {\"type\":\"Feature\",\"properties\":{\"name\":\"Plot A\",\"adm_level\":100},\"geometry\":{...}} |
geom | No | WKT geometry (POINT, LINESTRING, POLYGON, MULTIPOLYGON). (Provide geom or lat+long.) | POLYGON((-60 -3,-60.1 -3,-60.1 -3.1,-60 -3.1,-60 -3)) |
ismarine | No | Flag to allow marine/offshore locations outside country polygons. | 1 |
lat | No | Latitude in decimal degrees (negative for south). (Provide geom or lat+long.) | -3.101 |
long | No | Longitude in decimal degrees (negative for west). (Provide geom or lat+long.) | -60.12 |
name | Yes | Translatable trait name. Accepts a plain string or a JSON map of language codes to names. | {"en":"Height","pt-br":"Altura"} |
notes | No | Free text or JSON notes field. | {\"expedition\":\"2024-01\",\"tag\":\"P1\"} |
parent | No | Parent id/name; for traits this is another trait export_name/id to define hierarchy. | woodDensity |
startx | No | Start X coordinate for subplot relative to parent plot. | 5.5 |
starty | No | Start Y coordinate for subplot relative to parent plot. | 10.0 |
x | No | X coordinate for plots or individual position. | 12.3 |
y | No | Y coordinate for plots or individual position. | 8.7 |
locations-validation (POST)
Validates coordinates against registered locations (POST).
| Parameter | Required | Description | Example |
|---|---|---|---|
latitude | Yes | Latitude in decimal degrees (negative for south). | -3.101 |
longitude | Yes | Longitude in decimal degrees (negative for west). | -60.12 |
measurements (POST)
Trait measurements (GET lists, POST creates/imports via ImportMeasurements job, PUT bulk updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
bibreference | No | Bibreference id or bibkey. | 34 or ducke1953 |
dataset | Yes | Dataset id/name where the measurement will be stored; falls back to authenticated user default dataset if omitted. | 3 or FOREST1 |
date | Yes | Measurement date; accepts YYYY-MM-DD, YYYY-MM, YYYY, or array/year-month-day fields (date_year/date_month/date_day). | 2024-05-10 or {\"year\":2024,\"month\":5} |
duplicated | No | Integer allowing repeated measurements on the same date/object; must be higher than existing duplicates. | 2 |
link_id | No | Required for LINK trait types: id of the linked object (e.g., Taxon id). (Required when trait type is Link.) | 55 |
location | No | Location id or name. | Parcela 25ha or 55 |
notes | No | Optional free text or JSON notes stored with the measurement. | {\"method\":\"caliper\"} |
object_id | Yes | Required. Id of the measured object (Individual, Location, Taxon, Voucher, Media). Alias: measured_id. | 4521 |
object_type | Yes | Required when not provided in header. Class basename or FQCN of the measured object (Individual, Location, Taxon, Voucher, Media). Alias: measured_type. | Individual |
parent_measurement | No | When trait depends on another measurement, provide the parent measurement id for the same object and date. | 3001 |
person | Yes | Person id, abbreviation, full name or email (supports lists with | or ;). | J.Silva|M.Costa |
trait_id | Yes | Required. Trait id or export_name to be measured (also accepts trait key “trait”). | DBH or 12 |
value | No | Input varies by trait type: QUANT_INTEGER (0) = integer number; QUANT_REAL (1) = decimal number with dot separator; CATEGORICAL or ORDINAL (2/4) = single category id or translated name; CATEGORICAL_MULTIPLE (3) = list of category ids/names separated by | ; or , (or an array); TEXT (5) = free text string; COLOR (6) = hex color like #A1B2C3 or #ABC; LINK (7) = send link_id pointing to the linked object (value may be blank or a numeric qualifier); SPECTRAL (8) = semicolon-separated numeric series whose length equals trait value_length; GENEBANK (9) = GenBank accession string (validated against NCBI). (Required unless trait type is Link.) | QUANT_REAL: 23.4 | CATEGORICAL: 15 or Dead | CATEGORICAL_MULTIPLE: 12;14 or Alternate;Opposite | SPECTRAL: 0.12;0.11;0.10 |
media (POST)
Media metadata (GET lists, POST creates, PUT updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
collector | No | Collector(s) id, abbreviation, name or email. Use | or ; to separate multiple people; first is main collector. | J.Silva|M.Costa |
dataset | No | Dataset id or acronym. | 3 or FOREST1 |
date | No | Date (YYYY-MM-DD) or incomplete date (e.g. 1888-05-NA) or array with year/month/day. | 2024-05-20 or {\"year\":1888,\"month\":5} |
filename | Yes | Exact media file name inside the ZIP when importing media. | IMG_0001.jpg |
latitude | No | Latitude in decimal degrees (negative for south). | -3.101 |
license | No | Public license code for media (CC0, CC-BY, CC-BY-SA, etc.). | CC-BY-SA |
location | No | Location id or name. | Parcela 25ha or 55 |
longitude | No | Longitude in decimal degrees (negative for west). | -60.12 |
notes | No | Free text or JSON notes field. | {\"expedition\":\"2024-01\",\"tag\":\"P1\"} |
object_id | Yes | Id of object the media belongs to (Individual, Location, Taxon, Voucher). | 4521 |
object_type | Yes | The object type the media belongs to, one of Individual, Location, Taxon, Voucher, or Media. | Individual |
project | No | Project id or acronym. | PDBFF or 2 |
tags | No | Tag ids or names list for media or filters (use | or ;). | flower|leaf |
title_en | No | Media title in English. | Leaf detail |
title_pt | No | Media title in Portuguese. | Detalhe da folha |
persons (POST)
People (GET lists, POST creates, PUT updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
abbreviation | No | Standard abbreviation for a person or biocollection. | Silva, J.B, Pilco, M.V. |
biocollection | No | Biocollection id, name or acronym. | INPA |
email | No | Email address. | user@example.org |
full_name | Yes | Person full name. | Joao Silva |
institution | No | Institution associated with a person. | INPA |
taxons (POST)
Taxonomic names (GET lists, POST creates).
| Parameter | Required | Description | Example |
|---|---|---|---|
author | No | Taxon author string for unpublished names. | Smith & Jones |
author_id | No | Person id/name/email for the author of an unpublished taxon. (Required for unpublished names (or use person).) | 25 or Pilco, M.V. |
bibreference | No | Bibreference id or bibkey. | 34 or ducke1953 |
gbif | No | GBIF nubKey for a taxon. | 28792 |
ipni | No | IPNI id for a taxon. | 123456-1 |
level | No | Taxon rank code or string. | 210 or species |
mobot | No | Tropicos id for a taxon. | 12345678 |
mycobank | No | MycoBank id for a taxon. | MB123456 |
name | Yes | Translatable trait name. Accepts a plain string or a JSON map of language codes to names. | {"en":"Height","pt-br":"Altura"} |
parent | No | Parent id/name; for traits this is another trait export_name/id to define hierarchy. (Required for unpublished names.) | woodDensity |
person | No | Person id, abbreviation, full name or email (supports lists with | or ;). (Required for unpublished names (or use author_id).) | J.Silva|M.Costa |
valid | No | When 1, return only valid taxon names. | 1 |
zoobank | No | ZooBank id for a taxon. | urn:lsid:zoobank.org:act:12345678 |
traits (POST)
Trait definitions (GET lists, POST creates).
| Parameter | Required | Description | Example |
|---|---|---|---|
bibreference | No | Bibreference id or bibkey. | 34 or ducke1953 |
categories | No | Trait categories JSON list with lang/rank/name/description. (Required for categorical and ordinal traits.) | [{\"lang\":\"en\",\"rank\":1,\"name\":\"small\"}] |
description | Yes | Translatable description text. Accepts a plain string or a JSON map of language codes to descriptions. | {"en":"Tree height at breast height","pt-br":"Altura da árvore à altura do peito"} |
export_name | Yes | Unique export name for trait. | treeDbh,plantHeight |
link_type | No | Class name for Link trait target (e.g. Taxon). (Required for Link traits.) | Taxon |
name | Yes | Translatable trait name. Accepts a plain string or a JSON map of language codes to names. | {"en":"Height","pt-br":"Altura"} |
objects | Yes | Trait target objects (comma separated). | Individual,Voucher |
parent | No | Parent trait id or export_name; when set, measurements of this trait must also include a measurement for the parent trait. | woodDensity |
range_max | No | Maximum allowed numeric value for quantitative traits. | 999.9 |
range_min | No | Minimum allowed numeric value for quantitative traits. | 0.01 |
tags | No | Tag ids or names list for media or filters (use | or ;). | flower|leaf |
type | Yes | Generic type parameter (trait type code or vernacular type such as use/generic/etimology). | use or 10 |
unit | No | (Required for quantitative traits.) | — |
value_length | No | Number of values for spectral trait types. (Required for spectral traits.) | 1024 |
wavenumber_max | No | Maximum wavenumber for spectral traits. (Required for spectral traits.) | 25000 |
wavenumber_min | No | Minimum wavenumber for spectral traits. (Required for spectral traits.) | 4000 |
vernaculars (POST)
Vernacular names (GET lists, POST creates).
| Parameter | Required | Description | Example |
|---|---|---|---|
citations | No | List of citations (text + bibreference) for vernaculars. | [{\"citation\":\"Silva 2020\",\"bibreference\":12}] |
individuals | No | List of individual ids/fullnames for vernacular links. | 12|23|45 |
language | Yes | Language id/code/name | en or 1 or english or portuguese |
name | Yes | Translatable trait name. Accepts a plain string or a JSON map of language codes to names. | {"en":"Height","pt-br":"Altura"} |
notes | No | Free text or JSON notes field. | {\"expedition\":\"2024-01\",\"tag\":\"P1\"} |
parent | No | Parent id/name; for traits this is another trait export_name/id to define hierarchy. | woodDensity |
taxons | No | List of taxon ids/names (vernacular links). | Euterpe edulis|Euterpe precatoria |
type | No | Generic type parameter (trait type code or vernacular type such as use/generic/etimology). | use or 10 |
vouchers (POST)
Voucher specimens (GET lists, POST creates, PUT updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
biocollection | Yes | Biocollection id, name or acronym. | INPA |
biocollection_number | No | Catalogue number/code inside the biocollection. | 12345 |
biocollection_type | No | Nomenclatural type code or name. | Holotype or 2 |
collector | No | Collector(s) id, abbreviation, name or email. Use | or ; to separate multiple people; first is main collector. | J.Silva|M.Costa |
dataset | No | Dataset id or acronym. | 3 or FOREST1 |
date | No | Date (YYYY-MM-DD) or incomplete date (e.g. 1888-05-NA) or array with year/month/day. | 2024-05-20 or {\"year\":1888,\"month\":5} |
individual | Yes | Individual id, uuid or organismID (fullname). | 4521 or 2ff0e884-3d33 |
notes | No | Free text or JSON notes field. | {\"expedition\":\"2024-01\",\"tag\":\"P1\"} |
number | No | Collector number/code (voucher/individual tag when different from individual). | 1234A |
datasets (POST)
Datasets and published dataset files (GET lists, POST creates via import job).
| Parameter | Required | Description | Example |
|---|---|---|---|
description | No | Translatable description text. Accepts a plain string or a JSON map of language codes to descriptions. (Required when privacy is 2 or 3.) | {"en":"Tree height at breast height","pt-br":"Altura da árvore à altura do peito"} |
license | No | Public license code for media (CC0, CC-BY, CC-BY-SA, etc.). (Required when privacy is 2 or 3.) | CC-BY-SA |
name | Yes | Short name or nickname for the dataset - make informative, shorter than title. | Morphometrics-Aniba |
privacy | Yes | (Accepted values: 0 (auth), 1 (project), 2 (registered), 3 (public).) | — |
project_id | No | (Required when privacy is 1 (project).) | — |
title | No | (Required when privacy is 2 or 3.) | — |
3.4 - Put data
Only the endpoints listed below can be updated using the API and only the listed PUT fields can be updated on each endpoint. Field values are as explained for the POST API endpoints, except that in all cases you must also provide the id of the record to be updated.
Quick links
individuals (PUT)
Individuals (GET lists, POST creates, PUT updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Numeric ID of the record to be updated (Provide id or individual_id.) | 12 |
collector | No | Collector(s) id, abbreviation, name or email. Use | or ; to separate multiple people; first is main collector. | J.Silva|M.Costa |
dataset | No | Dataset id or acronym. | 3 or FOREST1 |
date | No | Date (YYYY-MM-DD) or incomplete date (e.g. 1888-05-NA) or array with year/month/day. | 2024-05-20 or {\"year\":1888,\"month\":5} |
identification_based_on_biocollection | No | Biocollection name/id used as reference for identification. | INPA |
identification_based_on_biocollection_number | No | CatalogNumber in the reference biocollection | 8765 |
identification_date | No | Identification date (full or incomplete). | 2023-06-NA |
identification_individual | No | ID/organismID of another individual that provides the taxonomic identification for the individual’s record | 3245 or REC-123 |
identification_notes | No | Notes for the identification. | Checked with microscope |
identifier | No | Person(s) responsible for identification; accept id, abbreviation, name or email; separate multiple with | or ;. | Costa, A.|Lima, B. or 1|2|3 or Adolpho Ducke|José Ramos|Paulo Apóstolo Costa Lima Assunção |
individual_id | No | Numeric ID of the record to be updated (Provide id or individual_id.) | 12 |
modifier | No | Identification modifier code/name (s.s.=1, s.l.=2, cf.=3, aff.=4, vel aff.=5). | 3 |
notes | No | Free text or JSON notes field. | {\"expedition\":\"2024-01\",\"tag\":\"P1\"} |
tag | No | Individual tag/number/code. | A-1234 |
taxon | No | Taxon id or canonical full name list. | Licaria cannela or 456,789 |
individual-locations (PUT)
Occurrences for individuals with multiple locations (GET lists, POST/PUT upserts).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Numeric ID of the record to be updated (Provide id or individual_location_id.) | 12 |
altitude | No | Elevation in meters. | 75 |
angle | No | Azimuth from reference point, in degrees. | 45 |
distance | No | Distance from reference point in meters. | 12.5 |
individual | No | Individual id, uuid or organismID (fullname). | 4521 or 2ff0e884-3d33 |
individual_location_id | No | Individual-location record id. (Provide id or individual_location_id.) | 44 |
latitude | No | Latitude in decimal degrees (negative for south). | -3.101 |
location | No | Location id or name. | Parcela 25ha or 55 |
location_date_time | No | Date or date+time for the occurrence/location event. | 2023-08-14 12:30:00 |
location_notes | No | Notes for the occurrence/location entry. | Near trail marker 10 |
longitude | No | Longitude in decimal degrees (negative for west). | -60.12 |
x | No | X coordinate for plots or individual position. | 12.3 |
y | No | Y coordinate for plots or individual position. | 8.7 |
locations (PUT)
Locations (GET lists, POST creates, PUT updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Numeric ID of the record to be updated (Provide id or location_id.) | 12 |
adm_level | No | Location administrative level code (e.g. 100=plot, 10=country). | 100 |
altitude | No | Elevation in meters. | 75 |
datum | No | Spatial datum/projection string. | EPSG:4326-WGS 84 |
geom | No | WKT geometry (POINT, LINESTRING, POLYGON, MULTIPOLYGON). | POLYGON((-60 -3,-60.1 -3,-60.1 -3.1,-60 -3.1,-60 -3)) |
ismarine | No | Flag to allow marine/offshore locations outside country polygons. | 1 |
lat | No | Latitude in decimal degrees (negative for south). | -3.101 |
location_id | No | Location id of the record to update. (Provide id or location_id.) | 44 |
long | No | Longitude in decimal degrees (negative for west). | -60.12 |
name | No | Translatable trait name. Accepts a plain string or a JSON map of language codes to names. | {"en":"Height","pt-br":"Altura"} |
notes | No | Free text or JSON notes field. | {\"expedition\":\"2024-01\",\"tag\":\"P1\"} |
parent | No | Parent id/name; for traits this is another trait export_name/id to define hierarchy. | woodDensity |
startx | No | Start X coordinate for subplot relative to parent plot. | 5.5 |
starty | No | Start Y coordinate for subplot relative to parent plot. | 10.0 |
x | No | X coordinate for plots or individual position. | 12.3 |
y | No | Y coordinate for plots or individual position. | 8.7 |
measurements (PUT)
Trait measurements (GET lists, POST creates/imports via ImportMeasurements job, PUT bulk updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Numeric ID of the record to be updated (Provide id or measurement_id.) | 12 |
bibreference | No | Bibreference id or bibkey. | 34 or ducke1953 |
dataset | No | Dataset id or acronym. | 3 or FOREST1 |
date | No | Date (YYYY-MM-DD) or incomplete date (e.g. 1888-05-NA) or array with year/month/day. | 2024-05-20 or {\"year\":1888,\"month\":5} |
duplicated | No | Sequential number to allow duplicate measurements for same trait+object+date. | 2 for a second measurement, number 3 for third, ... |
link_id | No | Linked object id when trait type is Link. | taxon id 55 |
location | No | Location id or name. | Parcela 25ha or 55 |
measurement_id | No | Measurement id of the record to update. (Provide id or measurement_id.) | 77 |
notes | No | Free text or JSON notes field. | {\"expedition\":\"2024-01\",\"tag\":\"P1\"} |
object_id | No | Id of the measured object (Individual, Location, Taxon, Voucher or Media). | 4521 |
object_type | No | Measured object type: Individual, Location, Taxon, Voucher, or Media. | Individual |
parent_measurement | No | Parent measurement ID for measurements whose variable depends on the measurement of another variable | 3001 |
person | No | Person id, abbreviation, full name or email (supports lists with | or ;). | J.Silva|M.Costa |
trait_id | No | Trait id or export_name for measurements. | 12 or DBH |
value | No | Measurement payload varies by trait type: QUANT_INTEGER (0) = integer number; QUANT_REAL (1) = decimal number with dot separator; CATEGORICAL or ORDINAL (2/4) = single category id or translated name; CATEGORICAL_MULTIPLE (3) = list of category ids/names separated by | ; or , (or an array); TEXT (5) = free text string; COLOR (6) = hex color like #A1B2C3 or #ABC; LINK (7) = send link_id pointing to the linked object (value may be blank or a numeric qualifier); SPECTRAL (8) = semicolon-separated numeric series whose length equals trait value_length; GENEBANK (9) = GenBank accession string (validated against NCBI). | QUANT_REAL: 23.4 | CATEGORICAL: 15 | CATEGORICAL_MULTIPLE: 12|14 | SPECTRAL: 0.12;0.11;0.10 |
media (PUT)
Media metadata (GET lists, POST creates, PUT updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Single id or comma-separated list to filter or target records. (Provide id, media_id or media_uuid.) | 1,2,3 |
collector | No | Collector(s) id, abbreviation, name or email. Use | or ; to separate multiple people; first is main collector. | J.Silva|M.Costa |
dataset | No | Dataset id or acronym. | 3 or FOREST1 |
date | No | Date (YYYY-MM-DD) or incomplete date (e.g. 1888-05-NA) or array with year/month/day. | 2024-05-20 or {\"year\":1888,\"month\":5} |
latitude | No | Latitude in decimal degrees (negative for south). | -3.101 |
license | No | Public license code for media (CC0, CC-BY, CC-BY-SA, etc.). | CC-BY-SA |
location | No | Location id or name. | Parcela 25ha or 55 |
longitude | No | Longitude in decimal degrees (negative for west). | -60.12 |
media_id | No | Media numeric id. (Provide id, media_id or media_uuid.) | 88 |
media_uuid | No | Media UUID. (Provide id, media_id or media_uuid.) | a3f0a4ac-6b5b-11ed-b8c0-0242ac120002 |
notes | No | Free text or JSON notes field. | {\"expedition\":\"2024-01\",\"tag\":\"P1\"} |
project | No | Project id or acronym. | PDBFF or 2 |
tags | No | Tag ids or names list for media or filters (use | or ;). | flower|leaf |
title_en | No | Media title in English. | Leaf detail |
title_pt | No | Media title in Portuguese. | Detalhe da folha |
persons (PUT)
People (GET lists, POST creates, PUT updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Numeric ID of the record to be updated (Provide id or person_id.) | 12 |
abbreviation | No | Standard abbreviation for a person or biocollection. | Silva, J.B, Pilco, M.V. |
biocollection | No | Biocollection id, name or acronym. | INPA |
email | No | Email address. | user@example.org |
full_name | No | Person full name. | Joao Silva |
institution | No | Institution associated with a person. | INPA |
person_id | No | Person id of the record to update. (Provide id or person_id.) | 12 |
vouchers (PUT)
Voucher specimens (GET lists, POST creates, PUT updates).
| Parameter | Required | Description | Example |
|---|---|---|---|
id | No | Numeric ID of the record to be updated (Provide id or voucher_id.) | 12 |
biocollection | No | Biocollection id, name or acronym. | INPA |
biocollection_number | No | Catalogue number/code inside the biocollection. | 12345 |
biocollection_type | No | Nomenclatural type code or name. | Holotype or 2 |
collector | No | Collector(s) id, abbreviation, name or email. Use | or ; to separate multiple people; first is main collector. | J.Silva|M.Costa |
dataset | No | Dataset id or acronym. | 3 or FOREST1 |
date | No | Date (YYYY-MM-DD) or incomplete date (e.g. 1888-05-NA) or array with year/month/day. | 2024-05-20 or {\"year\":1888,\"month\":5} |
individual | No | Individual id, uuid or organismID (fullname). | 4521 or 2ff0e884-3d33 |
notes | No | Free text or JSON notes field. | {\"expedition\":\"2024-01\",\"tag\":\"P1\"} |
number | No | Collector number/code (voucher/individual tag when different from individual). | 1234A |
voucher_id | No | Voucher id of the record to update. (Provide id or voucher_id.) | 55 |
4 - Concepts
If you want help development, read carefully the OpenDataBio data model concept before start collaborating.
To facilitate the understanding of the concepts, as include many tables and complex relationships, the OpenDataBio data model is divided in the four categories listed below.
4.1 - Core Objects
Core objects are: Location, Voucher, Individual and Taxon. These entities are considered “Core” because they may have Measurements, i.e. you may register values for any custom Trait.
The Individual object refer to Individual organism that have been observed once (an occurrence) or has been tagged for monitoring, such as tree in a permanent plot, a banded bird, a radio-tracked bat. Individuals may have one or more Vouchersin a BioCollection, and one or multiple Locations, and will have a taxonomic Identification. Any attribute measured or taken for individual organism may be associated with this object through the Measurement Model model.
The Voucherobject is for records of specimens from Individuals deposited in a Biological Collection. The taxonomic Identification and the Location of a Voucher is that of the Individual it belongs to. Measurements may be linked to a Voucher when you want to explicitly register the data to that particular sample (e.g. morphological measurements; a molecular marker from an extraction of a sample in a tissue collection). Otherwise you could just record the Measurement for the Individual the Voucher belongs to. The voucher model is also available as special type of Trait, the
LinkType, making it possible to record counts for the voucher’s Taxon at a particular Location.The Location object contains spatial geometries, like points and polygons, and include
plotsandtransectsas special cases. An Individual may have one location (e.g. a plant) or more locations (e.g. a monitored animal). Plots and Transect locations may be registered as a spatial geometry or only point geometry, and may have Cartesian dimensions (meters) registered. Individuals may also have Cartesian positions (X and Y or Angle and Distance) relative to their Location, allowing to account for traditional mapping of individuals in sampling units. Ecological relevant measurements, such as soil or climate data are examples of measurements that may be linked to locations Measurement.The Taxon object in addition to its use for the Identification of Individuals, may receive Measurements, allowing the organization of secondary, published data, or any kind of information linked to a Taxonomic name. A BibReference may be included to indicate the data source. Moreover, the Taxon model is available as special type of Trait, the
LinkType, making it possible to record counts for Taxons at a particular Location.
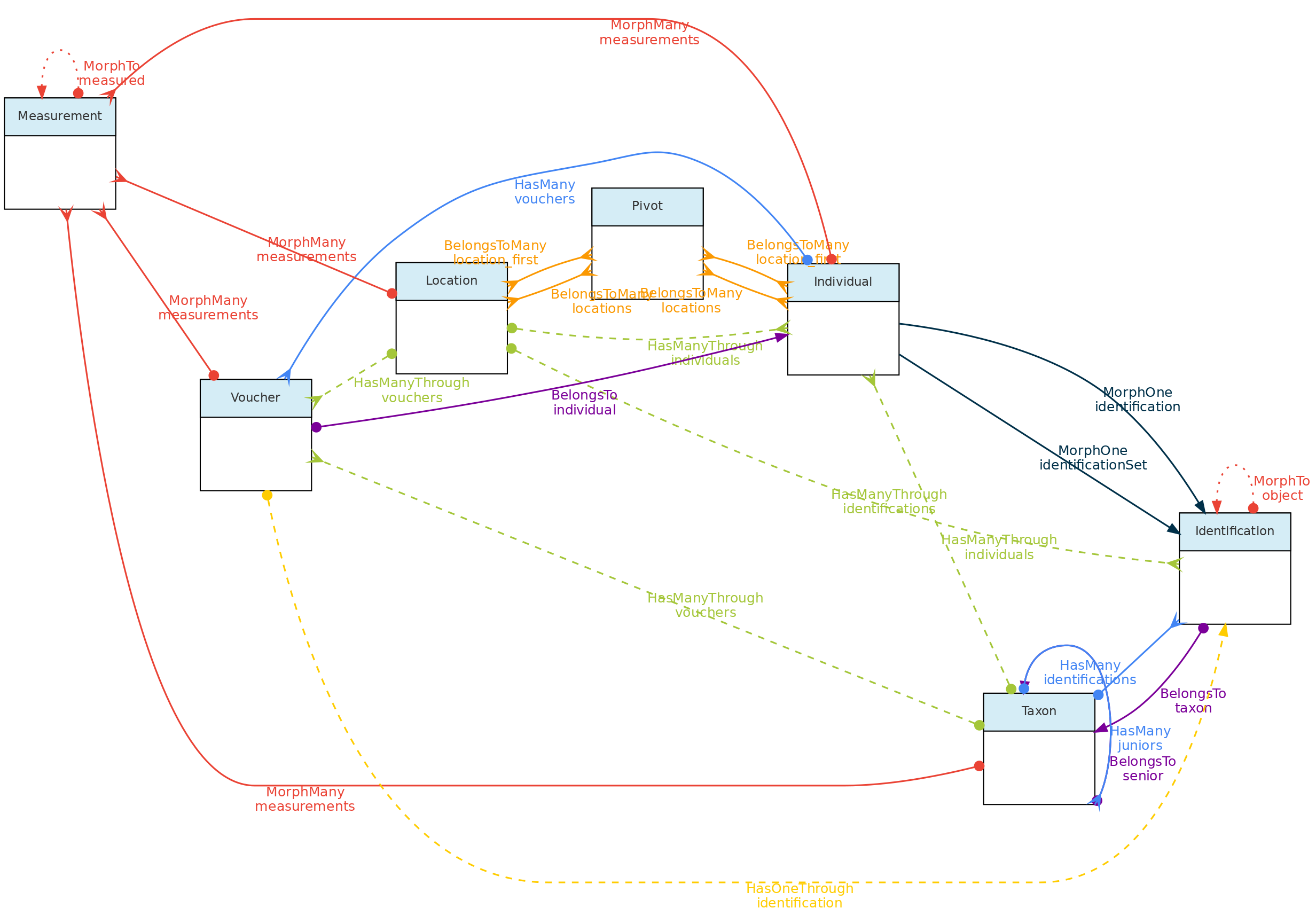
This figure show the relationships among the Core objects and with the Measurement Model. The Identification Model is also included for clarity. Solid links are direct relationships, while dashed links are indirect relationships (e.g. Taxons has many Vouchers through Individuals, and have many Individuals through identifications). The red solid lines link the Core objects with the Measurement model through polymorphic relationships. The dotted lines on the Measurement model just allow access to the measured core-object and to the models of link type traits.
Location Model
The Locations table stores data representing real world locations. They may be countries, cities, conservation units, or any spatial polygon, point or linestring on the surface of Earth. These objects are hierarchical and have a parent-child relationship implemented using the Nested Set Model for hierarchical data of the Laravel library Baum and facilite both validation and queries.
Special location types are plots and transects, which together with point locations allow different sampling methods used in biodiversity studies. These Location types may also be linked a parent location and in addition also to three additional types of location that may span different administrative boundaries, such as Conservation Units, Indigenous Territories and any Environmental layer representing vegetation classes, soil classes, etc…with defined spatial geometries.
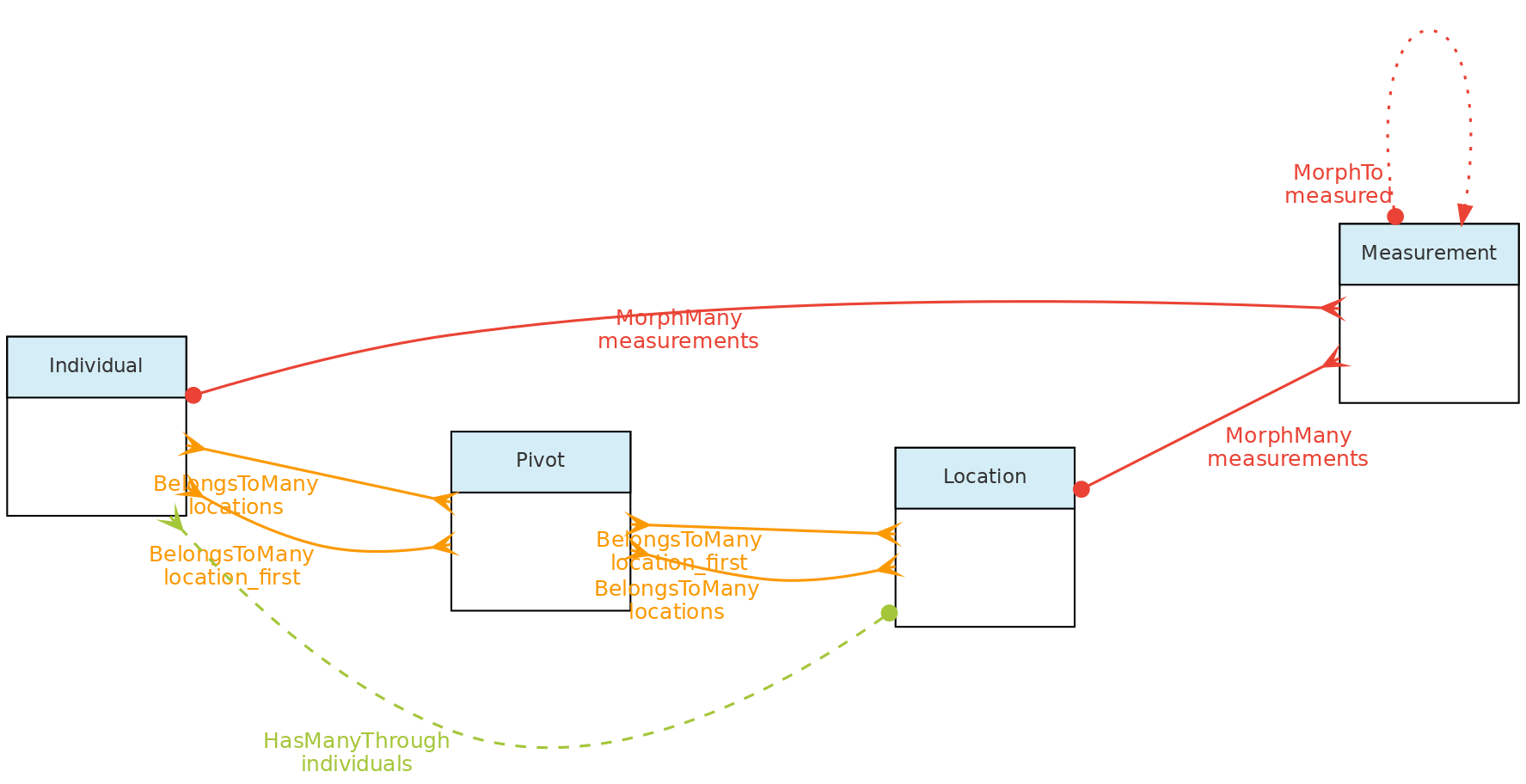 This figure shows the relationships of the
This figure shows the relationships of the Location model throught the methods implemented in the shown classes. The pivot table linking Location to Individual allow an individual to have multiple locations and each location for the individual to have specific attributes like date_time, altitude, relative_position and notes.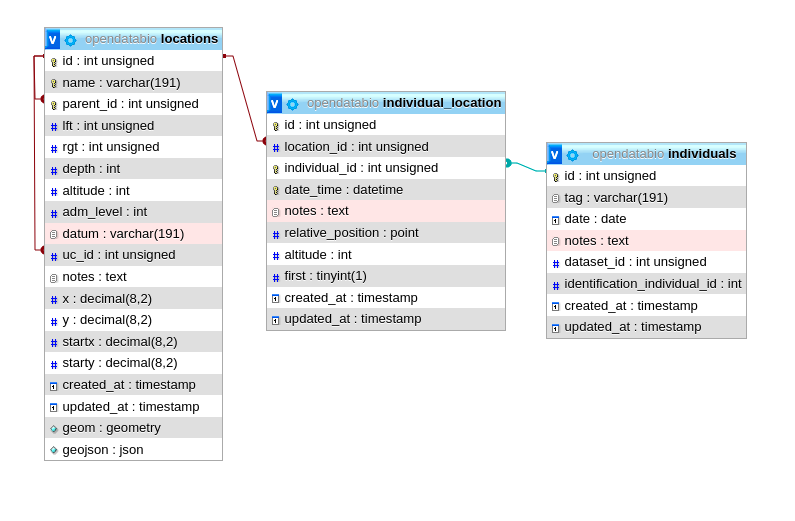 The same tables related with the Location model with the direct and non-polymoprhic relationships indicated.
The same tables related with the Location model with the direct and non-polymoprhic relationships indicated.
Location Table Columns
- Columns
parent_idtogether withrgt,lftanddephare used to define the Nested Set Model to query ancestors and descendants in a fast way. Onlyparent_idis specified by the user, the other columns are calculated by the Baum library trait from the id+parent_id values that define the hierarchy. The same hierarchical model is used for the Taxon Model, but for Locations there is a spatial constraint, i.e. a children must fall within a parent geometry. - The
adm_levelcolumn indicate the administrative level, or type, of a location. By default, the followingadm_levelare configured in OpenDataBio:2for country,3for first division within country (province, state),4for second division (e.g. municipality),… up toadm_level=10as administrative areas (country code is 2 to allow standardization with OpenStreeMaps, which is recommended to follow if your installation will include data from different countries). The administrative levels may be configured in an OpenDataBio before importing any data to the database, see the installation guide for details on that.99is the code for Conservation Units - a conservation unit is alocationthat may be linked to multiple other locations (any location may belong to a single UC). Thus, one Location may have as parent a city and as uc_id the conservation unit where it belongs.98is the code for Indigenous Territories - same properties as Conservation Units, but treated separately only because some CUs and TIs may largely overlap as is the case the Amazon region97ise the code for Environmental layers - same properties as Conservation Units and Indigenous Territories, i.e., may be linked as additional location to any Point, Plot or Transect, and thehence, their related individuals. Store polygons and multipolygon geometries representing environmental classes, such as vegetation units, biomes, soil classes, etc…100is the code forplotsand subplots - plot locations may be registered with Point or with a Polygon geometry, and must also have an associated Cartesian dimensions in meters. If it is a point location, the geometry is defined by ODB from the dimensions with NorthEast orientation from the point informed. Cartesian dimensions of a plot location can also be combined with cartesian positions of subplots (i.e. a plot location whose parent is also a plot location) and/or of individuals within such plots, allowing individuals and subplots to be mapped within aplotsubplot location without geometry specifications. In other words, if the spatial geometry of the plot is unknown, it may have as geometry a single GPS point rather than a polygon, plus itsxandydimensions. A subplot is location plot inside a location plot and must consist of a point marking the start of the subplot plus its X and Y cartesian dimensions. If the geometry of the start of the subplot is unknown, it may be stored as a relative position to parent plot using thestartxandstarty.101for transects - like plots, transects may be registered having a LineString geometry or simply a single Latitude and Longitude coordinates and a dimension. Thexcartesian dimension for transects represent the length in meters and is used to create a linestring (North oriented) when only a point is informed. Theydimension is used to validate individuals as belonging to transect location, and represents the maximum distance from the line that and individual must fall to be detected in that location.999for ‘POINT’ locations like GPS waypoints - this is for registration of any point in space
- Column
datummay record the geometry datum property, if known. If left blank, the location is considered to be stored using WGS84 datum. However, there is no built-in conversor from other types of data, so the maps displayed may be incorrect if different datum’s are used. Strongly recommended to project data as WSG84 for standardization. - Column
geomstores the location geometry in the database, allowing spatial queries in SQL language, such asparent autodetection. The geometry of a location may bePOINT,POLYGON,MULTIPOLYGONorLINESTRINGand must be formatted using Well-Known-Text geometry representation of the location. When a POLYGON is informed, the first point within the geometry string is privileged, i.e. it may be used as a reference for relative markings. For example, such point will be the reference for thestartxandstartycolumns of a subplot location. So forplotandtransectgeometries, it matters which point is listed first in the WKT geometry
Data access Full users may register new locations, edit locations details and remove locations records that have no associated data. Locations have open access!
Individual Model
The Individual object represents a record for an individual organism. It may be a single time-space occurrence of an animal, plant or fungi, or an individual monitored through time, such as a plant in a permanent forest plot, or an animal in capture-recapture or radio-tracking experiment.
An Individual may have one or more Vouchersrepresenting physical samples of the individual stored in one or more Biological Collection and it may have one or more Locations, representing the place or places where the individual has been recorded.
Individual objects may also have a self taxonomic Identification or its taxonomic identity may depend on that of another individual (non-self identification). The Individual identification is inherited by all the Vouchers registered for the Individual. Hence Vouchers do not have their separate identification.
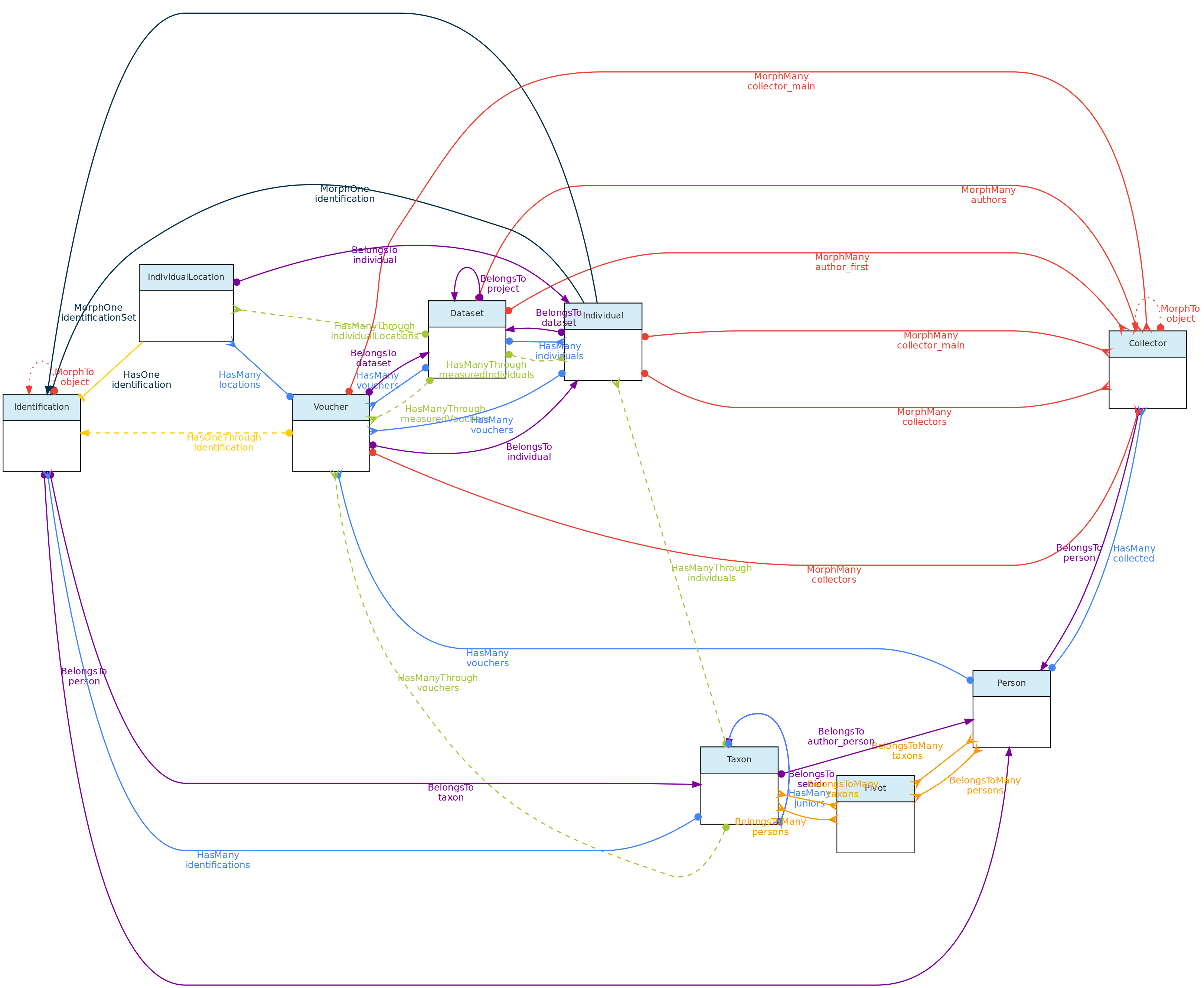 This figure shows the Individual Model and the models it relates to, except the Measurement and Location models, as their relationships with Individuals is shown elsewhere in this page. Lines linking models indicate the
This figure shows the Individual Model and the models it relates to, except the Measurement and Location models, as their relationships with Individuals is shown elsewhere in this page. Lines linking models indicate the methods or functions implemented in the classes to access the relationships. Dashed lines indicate indirect relationships and the colors the different types of Laravel Eloquent methods.
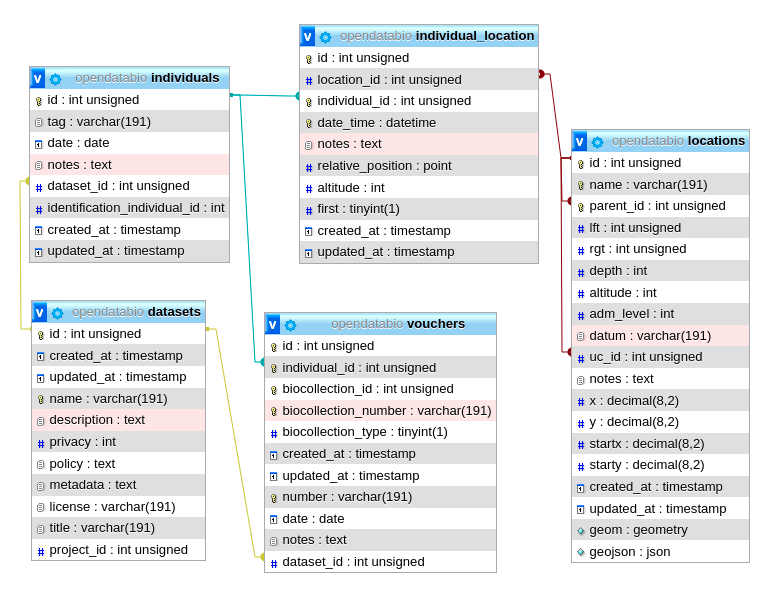 The Individual model direct and non-polymoprhic relationships.
The Individual model direct and non-polymoprhic relationships.
Individual Table Columns
- A Individual record must specify at least one Location where it was registered, the
dateof registration, the local identifiertag, and thecollectorsof the record, and thedataset_idthe individual belongs to. - The Location may be any location registered, regardless of level, allowing to store historical records whose georeferencing is just an administrative location. Individual locations are stored in the
individual_locationpivot table, having columnsdate_time,altitude,notesandrelative_positionfor the individual location records. - The column
relative_positionstores the Cartesian coordinates of the Individual in relation to its Location. This is only for individuals located in locations of typeplot,transectorpoint. For example, a Plot location with dimensions 100x100 meters (1ha) may have an Individual withrelative position=POINT(50 50), which will place the individual in the center of the location (this is shown graphically in the web-interface), as is defined by thexandycoordinates of the individual. If the location is a subplot, then the position within the parent plot may also be calculated (this was designed with ForestGeo plots in mind and is a column in the Individual GET API. If the location is a POINT, the relative_position may be informed asangle(= azimuth) anddistance, attributes frequently measured in sampling methods. If the location is a TRANSECT, the relative_position places the individual in relation to the linestring, thexbeing the distance along the transect from the first point, and theythe perpendicular distance where the individual is located, also accounting for some sampling methods; - The
datefield in the Individual, Voucher, Measurement and Identification models may be an Incomplete Date, i.e., only the year or year+month may be recorded. - The Collector table represents collectors for an Individual or Voucher, and is linked with the Person Model. The collector table has a polymorphic relationship with the Voucher and Individual objects, defined by columns
object_idandobject_type, allowing multiple collectors for each individual or voucher record. The main_collector indicated is just the first collector listed for these entities. - The
tagfield is a user code or identifier for the Individual. It may be the number written on the aluminum tag of a tree in a forest plot, the number of a bird-band, or thecollector numberof a specimen. The combination ofmain_collector+tag+first_locationis constrained to be unique in OpenDataBio. - The taxonomic identification of an Individual may be defined in two ways:
- for self identifications an Identification record is created in the identifications table, and the column
identification_individual_idis filled with the Individual ownid - for non-self identifications, the id of the Individual having the actual Identification is stored in column
identification_individual_id. - Hence, the Individual class contain two methods to relate to the Identification model: one that sets self identifications and another that retrieves the actual taxonomic identifications by using column
identification_individual_id.
- for self identifications an Identification record is created in the identifications table, and the column
- Individuals may have one or more Vouchersdeposited in a Biocollection.
Data access Individuals belong to Datasets, so Dataset access policy apply to the individuals in it. Only project collaborators and administrators may insert or edit individuals in a dataset, even if dataset is of public access.
Taxon Model
The general idea behind the Taxon model is to present tools for easily incorporating valid taxonomic names from Online Taxonomic Repositories (currently Tropicos.org and GBIF are implemented), but allowing for the inclusion of names that are not considered valid, either because they are still unpublished (e.g. a morphotype), or the user disagrees with published synonymia, or the user wants to have all synonyms registered as invalid taxons in the system. Moreover, it allows one to define a custom clade level for taxons, allowing one to store, in addition to taxonomic rank categories, any node of the tree of life. Any registered Taxon can be used in Individual identifications and Measurements may be linked to taxonomic names.
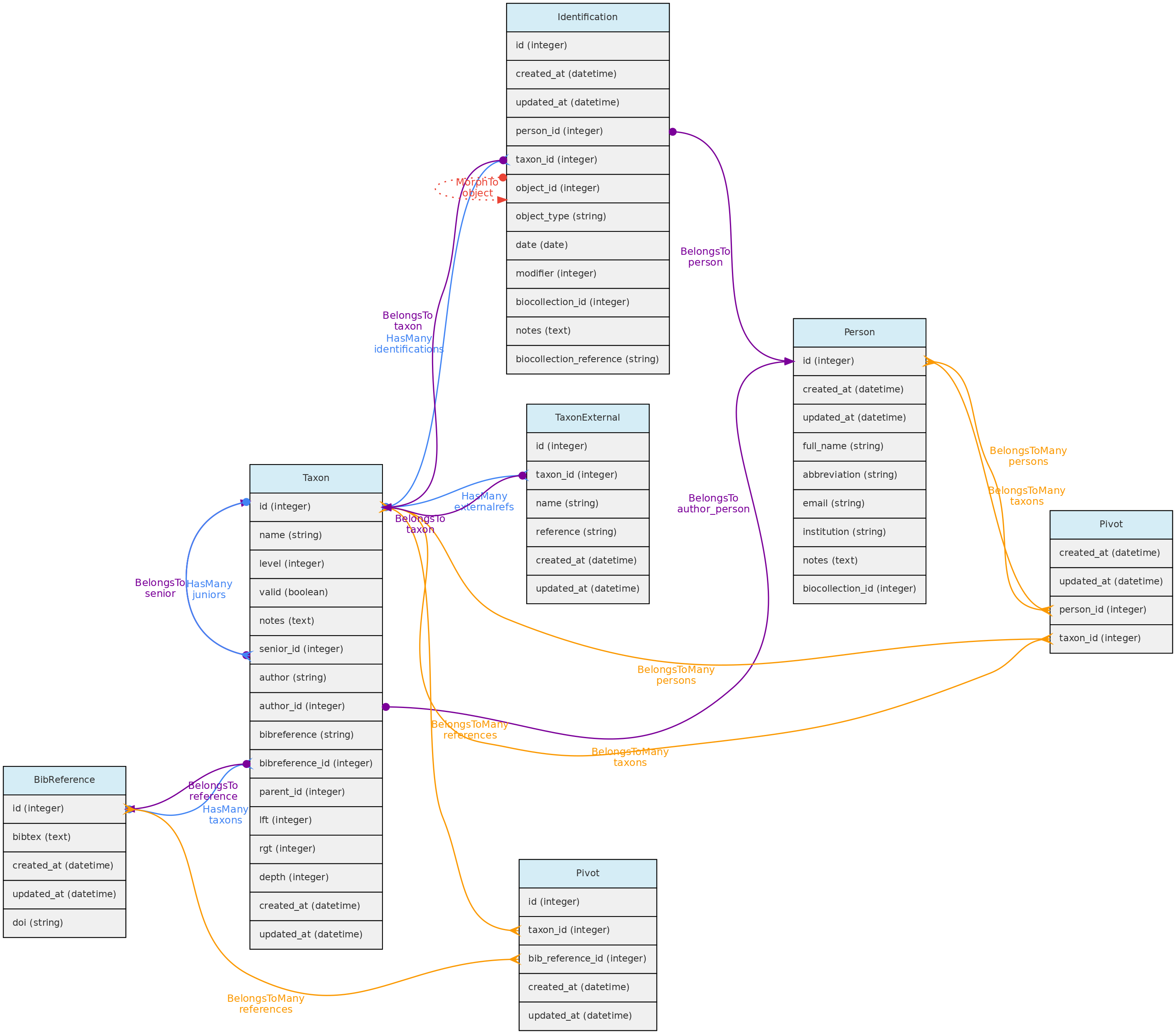 Taxon model and its relationships. Lines linking tables indicate the
Taxon model and its relationships. Lines linking tables indicate the methods implemented in the shown classes, with colors indicating different Eloquent relationships
Taxon table explained
- Like, Locations, the Taxon model has a parent-child relationship, implemented using the Nested Set Model for hierarchical data of the Laravel library Baum that allows to query ancestors and descendants. Hence, columns
rgt,lftanddephof the taxon table are automatically filled by this library upon data insertion or update. - For both, Taxon
authorand Taxonbibreferencethere are two options:- For published names, the string authorship retrieved by the external taxon APIs will be placed in the
author=stringcolumn. For unpublished names, author is a Person and will be stored in theauthor_idcolumn. - Only published names may have relation to BibReferences. The
bibreferencestring field of the Taxon table stores the strings retrieved through the external APIs, while thebibreference_idlinks to a BibReference object. These are used to store the Publication where the Taxon Name is described and may be entered in both formats. - In addition, a Taxon record may also have many other BibReferences through a pivot table (
taxons_bibreference), permitting to link any number of bibliographic references to a Taxon name.
- For published names, the string authorship retrieved by the external taxon APIs will be placed in the
- Column
levelrepresents the taxonomic rank (such as order, genera, etc). It is numerically coded and standardized following the IAPT general rules, but should accommodate also animal related taxon level categories. See the available codes in the Taxon API for the list of codes. - Column
parent_idindicates the parent of the taxon, which may be several levels above it. The parent level should be strictly higher than the taxon level, but you do not need to follow the full hierarchy. It is possible to register a taxon without parents, for example, an unpublished morphotype for which both genera and family are unknown may have anorderas parent. - Names for the taxonomic ranks are translated according to the system defined
localethat also translates the web interface (currently only Portuguese and English implemented). - The
namefield of the taxon table contain only the specific part of name (in case of species, the specific epithet), but the insertion and display of taxons through the API or webinterface should be done with the fullname combination. - It is possible to include synonyms in the Taxon table. To do so, one must fill in the
seniorrelationship, which is the id of the accepted (valid) name for aninvalidTaxon. Ifsenior_idis filled, then the taxon is ajuniorsynonym and must be flagged as invalid. - When inserting a new published taxon, only the
nameis required. The name will be validated and the author, reference and synonyms will be retrieved using the following API services:- GBIF BackBone Taxonomy - this will be the first check, from which links to Tropicos and IPNI may also be retrieved if registering a plant name.
- Tropicos - if not found on GBIF, ODB will search the name on the Missouri Botanical Garden nomenclature database.
- IPNI - the International Individual Names Index is another database used to validate individual names (Temporarily disabled)
- MycoBank - used to validate a name if not found by the Tropicos nor IPNI apis, and used to validate names for Fungi. Temporarily disabled
- ZOOBANK - when GBIF, Tropicos, IPNI and MycoBank fails to find a name, then the name is tested against the ZOOBANK api, which validates animal names. Does not provide taxon publication, however.
- If a Taxon name is found in the Nomenclatural databases, the respective ID of the repository is stored in the
taxon_externaltables, creating a link between the OpenDataBio taxon record and the external nomenclatural database. - A Person may be defined as one or more taxon specialist through a pivot table. So, a Taxon object may have many taxonomic specialist registered in OpenDataBio.
Data access: Full users are able to register a new taxon and edit existing records if they have not been used for Identification of Measurements. Currently it is impossible to remove a taxon from the database. Taxon list have public access.
Voucher Model
The Voucher model is used to store records of specimens or samples from Individuals deposited in Biological Collections. Therefore, the only mandatory information required to register a Voucher are individual, biocollection and whether the specimen is a nomenclature type (which defaults to non-type if not informed).
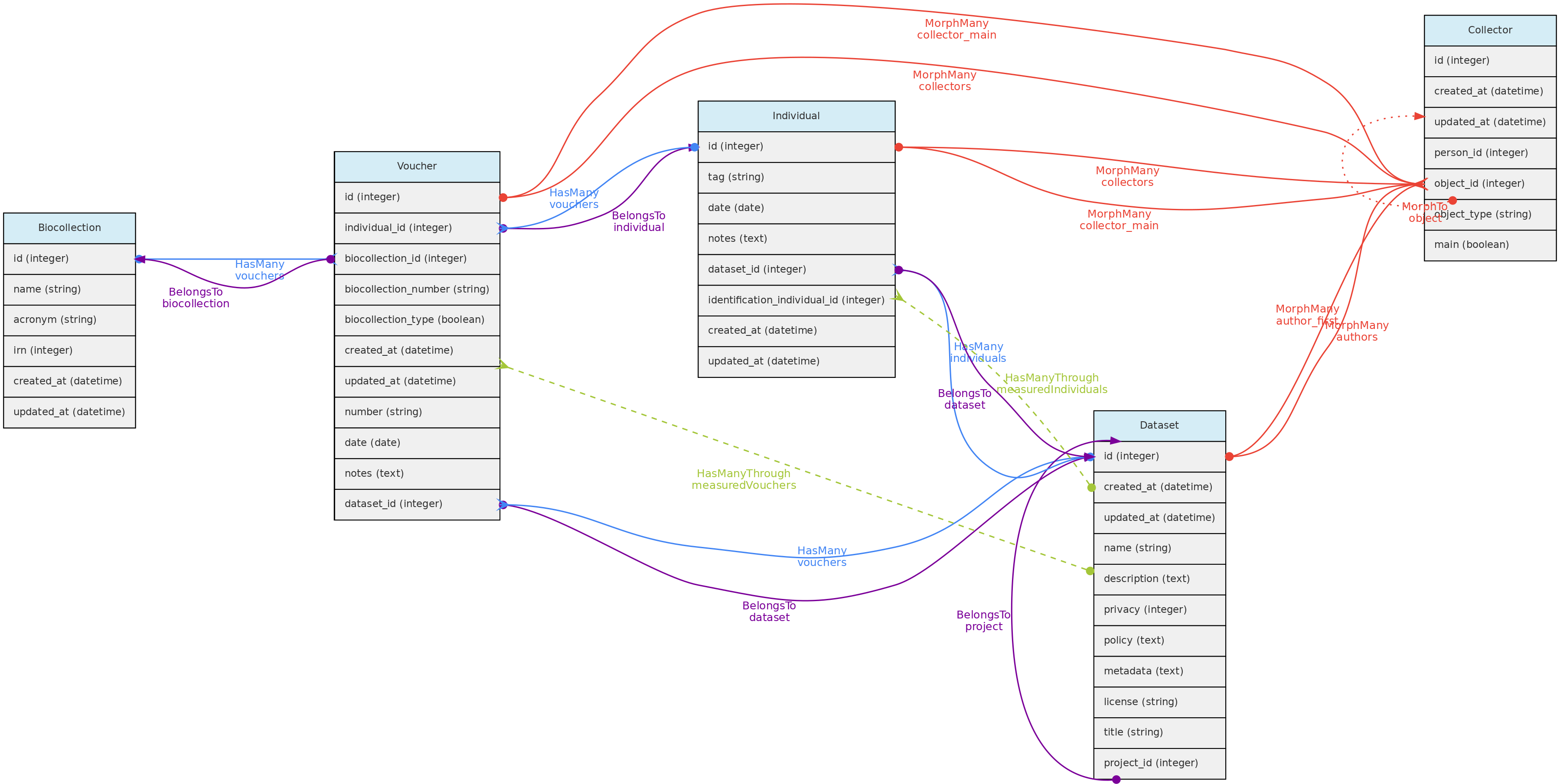 Voucher model and its relationships. Lines linking tables indicate the
Voucher model and its relationships. Lines linking tables indicate the methods implemented in the shown models, with colors indicating different Eloquent relationships. Not that Identification nor Location are show because Vouchers do not have their own records for these two models, they are just inherited from the Individual the Voucher belongs to
Vouchers table explained
- A Voucher belongs to an Individual and a Biocollection, so the
individual_idand thebiocollection_idare mandatory in this table; biocollection_numberis the alpha-numeric code of the Voucher in the BioCollection, it may be ’null’ for users that just want to indicate that a registered Individual have Vouchers in a particular Bicollection, or to registered Vouchers for biocollections that do not have an identifier code;biocollection_type- is a numeric code that specify whether the Voucher in the BioCollection is a nomenclatural type. Defaults to0(Not a Type);1for just ‘Type’, a generic form, and other numbers for other nomenclature type names (see the API Vouchers Endpoint for a full list of options).collectors, one or multiple, are optional for Vouchers, required only if they are different from the Individual collectors. Otherwise the Individual collectors are inherited by the Voucher. Like for Individuals, these are implemented through a polymorphic relationship with the collectors table and the first collector is the main_collector for the voucher, i.e. the one that relates tonumber.number, this is the collector number, but like collectors, should only be filled if different from the Individual’stagvalue. Hence,collectors,numberanddateare useful for registering Vouchers for Individuals that have Vouchers collected at different times by different people.datefield in the Individual and Voucher models may be an incomplete date. Only required if different from that of the Individual the Voucher belongs to.dataset_idthe Voucher belongs to a Dataset, which controls the access policy;notesany text annotation for the Voucher.- The Voucher model interacts with the BibReference model, permitting to link multiple citations to Vouchers. This is done with a pivot
voucher_bibreferencetable.
Data access Vouchers belong to Datasets, so Dataset access policy apply to the Vouchers in it. Vouchers may have a different Project than their Individuals. If the Voucher dataset policy is open access and that of the Individual project is not, then access to voucher data will be incomplete, so Voucher’s dataset should have the same or less restricted access policy than the Individual dataset. Only Dataset collaborators and administrators may insert or edit vouchers in a dataset, even if the dataset is of public access.
4.2 - Trait Objects
Measurement Model
The Measurements table stores the values for traits measured for core objects. Its relationship with the core objects is defined by a polymorphic relationship using columns measured_id and measured_type. These MorphTo relations are illustrated and explained in the core objects page.
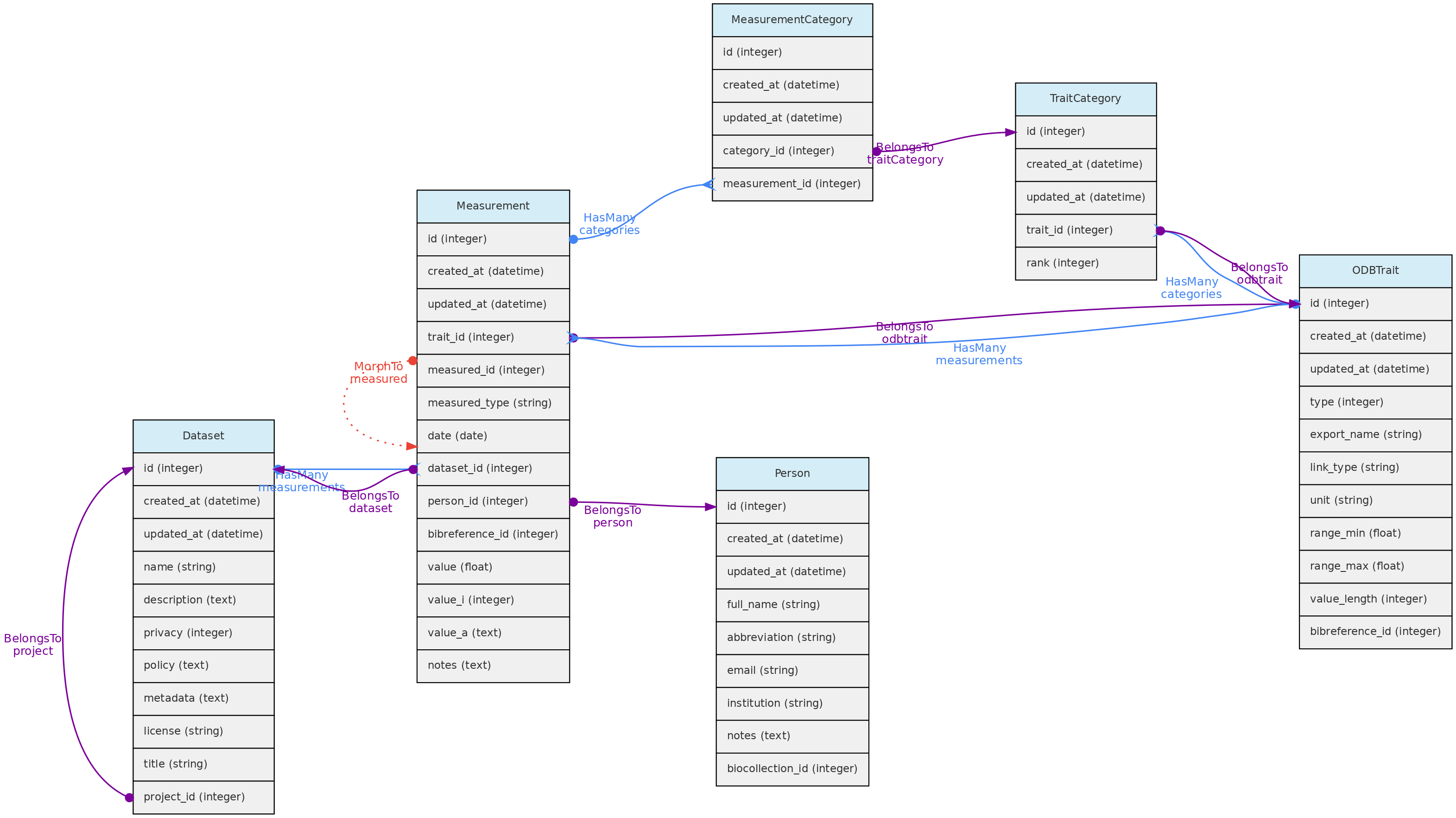
- Measurements must belong to a Dataset - column
dataset_id, which controls measurement access policy - A Person must be indicated as a measurer (
person_id); - The
bibreference_idcolumn may be used to link measurements extracted from publications to its Bibreference source; - The value for the measured trait (
trait_id) will be stored in different columns, depending on trait type:value- this float column will store values for Quantitative Real traits;value_i- this integer column will store values for Quantitative Integer traits; and is an optional field for Link type traits, allowing for example to store counts for a species (a Taxon Link trait) in a location.value_a- this text column will store values for Text, Color and Spectral trait types.
- Values for Categorical and Ordinal traits are stored in the
measurement_categorytable, which links measurements to trait categories. date- measurement date is mandatory in all cases
Data access Measurements belong to Datasets, so Dataset access policy apply to the measurements in it. Only dataset collaborators and administrators may insert or edit measurements in a dataset, even if the dataset is of public access.
Trait Model
The ODBTrait table represents user defined variables to collect Measurements for one of the core object, either Individual, Voucher, Location or Taxon.
These custom traits give enormous flexibility to users to register their variables of interest. Clearly, such flexibility has a cost in data standardization, as the same variable may be registered as different Traits in any OpenDataBio installation. To minimize redundancy in trait ontology, users creating traits are warned about this issue and a list of similar traits is presented in case found by trait name comparison.
Traits have editing restrictions to avoid data loss or unintended data meaning change. So, although the Trait list is available to all users, trait definitions may not be changed if somebody else also used the trait for storing measurements.
Traits are translatable entities, so their name and description values can be stored in multiple languages (see User Translations. This is placed in the user_translations table through a polymorphic relationship.
The Trait definition should be as specific as needed. The measurement of tree heights using direct measurement or a clinometer, for example, may not be easily converted from each other, and should be stored in different Traits. Thus, it is strongly recommended that the Trait definition field include information such as measurement instrument and other metadata that allows other users to understand whether they can use your trait or create a new one.
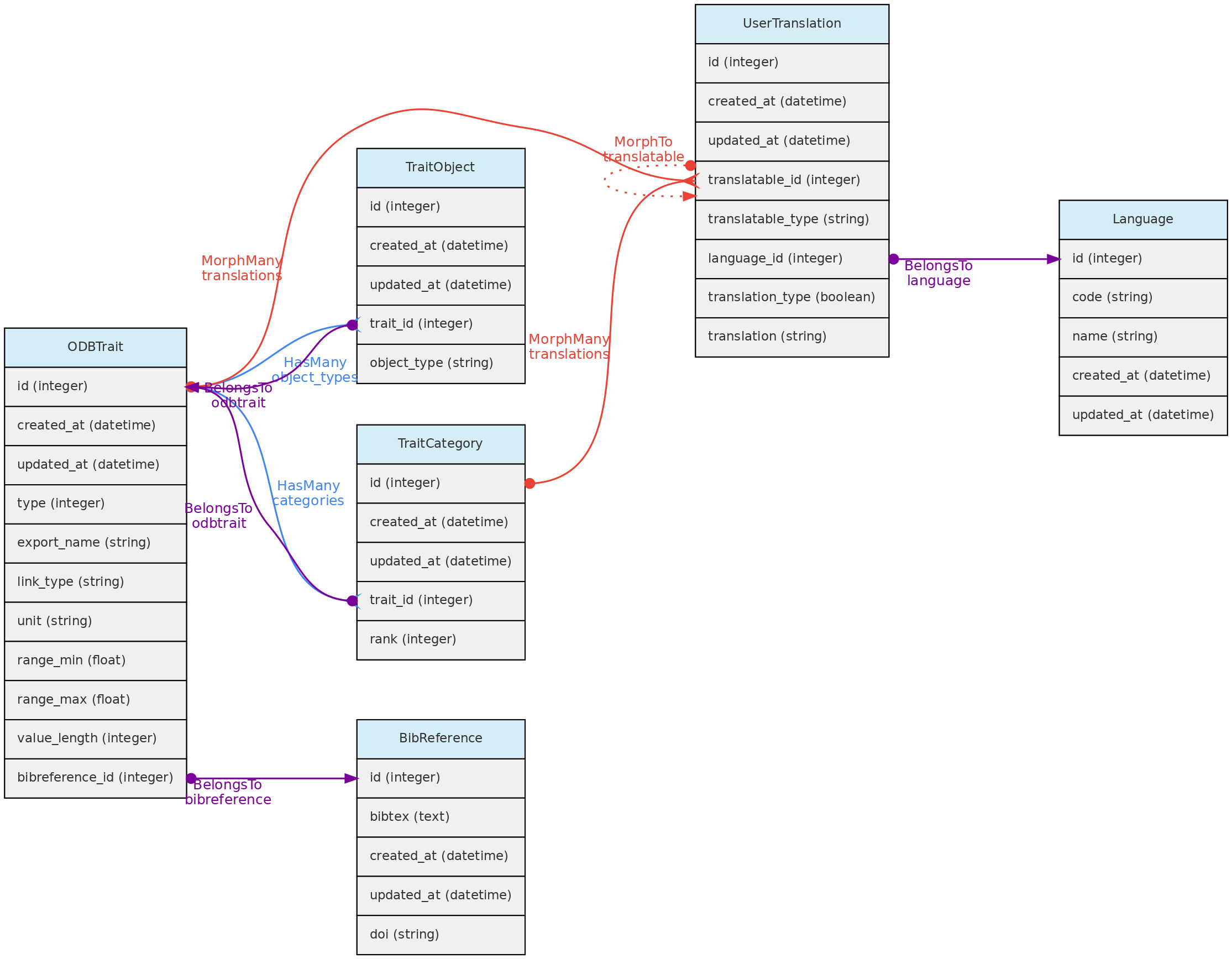
- The Trait definition must include an
export_namefor the trait, which will be used during data exports and are more easily used in trait selection inputs in the web-interface. Export names must be unique and have no translation. Short and camelCase or PascalCase export names are recommended. - The following trait types are available:
- Quantitative real - for real numbers;
- Quantitative integer - for counts;
- Categorical - for one selectable categories;
- Categorical multiple - for many selectable categories;
- Categorical ordinal - for one selectable ordered categories (semi-quantitative data);
- Text - for any text value;
- Color - for any color value, specified by the hexadecimal color code, allowing renderizations of the actual color.
- Link - this is a special trait type in OpenDataBio to link to database object. Currently, only link to Taxons and Voucher are allowed as a link type traits. Use ex: if you want to store species counts conducted in a location, you may create a Taxon link type Trait or a Voucher link type Trait if the taxon has vouchers. A measurement for such trait will have an optional
valuefield to store the counts. This trait type may also be used to specify the host of a parasite, or the number of predator insects. - Spectral - this is designed to accommodate Spectral data, composed of multiple absorbance or reflectance values for different wavenumbers.
- GenBank - this stores GenBank accessions numbers allowing to retrieve molecular data linked to individuals or vouchers stored in the database through the GenBank API Service.
- The Traits table contains fields that allow measurement value validation, depending on trait type:
range_maxandrange_min- if defined for Quantitative traits, measurements will have to fit the specified range;value_length- mandatory for Spectral Traits only, validate the length (number of values) of a spectral measurement;link_type- if trait is Link type, the measurementvalue_imust be an id of the link type object;- Color traits are validated in the measurement creation process and must conform to a color hexadecimal code. A color picker is presented in the web interface for measurement insertion and edition;
- Categorical and ordinal traits will be validated for the registered categories when importing measurements through the API;
- Column
unitdefines the measurement unit for the Trait. There is no way to prevent measurements values imported with a distinct unit. Quantitative traits requiredunitdefinition. - Column
bibreference_idis the key of a single BibReference that may be linked to trait definition. - The
trait_objectstable stores the type of core object (Taxon, Location, Voucher) that the trait can have a measurement for;
Data access A Trait name, definition, unit and categories may not be updated or removed if there is any measurement of this trait registered in the database. The only exceptions are: (a) it is allowed to add new categories to categorical (not ordinal) traits; (b) the user updating the trait is the only Person that has measurements for the trait; (c) the user updating the trait is an Admin of all the datasets having measurements using trait.
Forms
A Form is an organized group of Traits, defined by a User in order to create a custom form that can be filled in for entering measurements through the web interface. A Form consists of a group of ordered Traits, which can be marked as “mandatory”. Related entities are the Report and the Filter.
This is still experimental and needs deeper testing
4.3 - Data Access Objects
Datasets control data access and represents a dynamic data publication, with a version defined by last edition date. Datasets may contain Measurements, Individuals, Vouchers and/or Media Files.
Projects are just groups of Datasets and Users, representing coohorts of users with common accessibility to datasets whose privacy are set to be controlled by a Project.
BioCollections - This model serves to create a reusable list of acronyms of Biological Collections to record Vouchers. However, you can optionally manage a collection of Vouchers and their Individuals, in parallel to the access control provided by Datasets. Control is only for editing and entering Voucher records. In this case, the BioCollection is managed by the system.
Projects and BioCollections must have at least one User defined as administrator, who has total control over the dataset or project, including granting the following roles to other users: administrator, collaborator or viewer:
- Collaborators are able to insert and edit objects, but are not able to delete records nor change the dataset or project configuration.
- Viewers have read-only access to the data that are not of open access.
- Only Full Users and SuperAdmins may be assigned as administrators or collaborators. Thus, if a user who was administrator or collaborator of a dataset is demoted to “Registered User”, she or he will become a viewer.
- Only Super-admins can enable a BioCollection to be administered by the system.
Biocollections
The Biocollection model has two functions: (1) to provide a list of acronyms for registering Vouchers of any Biological Collection; (2) to manage the data of Biological Collections, facilitating the registration of new data (any user enters their data using the validations carried out by the software and requests the collection’s curators through the interface to register the data, which is done by authorized users for the BioCollection. Upon data registration, the BioCollection controls the editing of the data of the Vouchers and the related Individuals. Option (2) needs to be implemented by a Super-Administrator user, who can enable a BioCollection to be administered by the system, implementing the ODBRequest request model so that users can request data, samples, or records and changes to the data.
The Biocollection object may be a formal Biocollection, such as those registered in the Index Herbariorum (http://sweetgum.nybg.org/science/ih/), or any other Biological Collection, formal or informal.
The Biocollection object also interacts with the Person model. When a Person is linked to an Biocollection it will be listed as a taxonomic specialist.
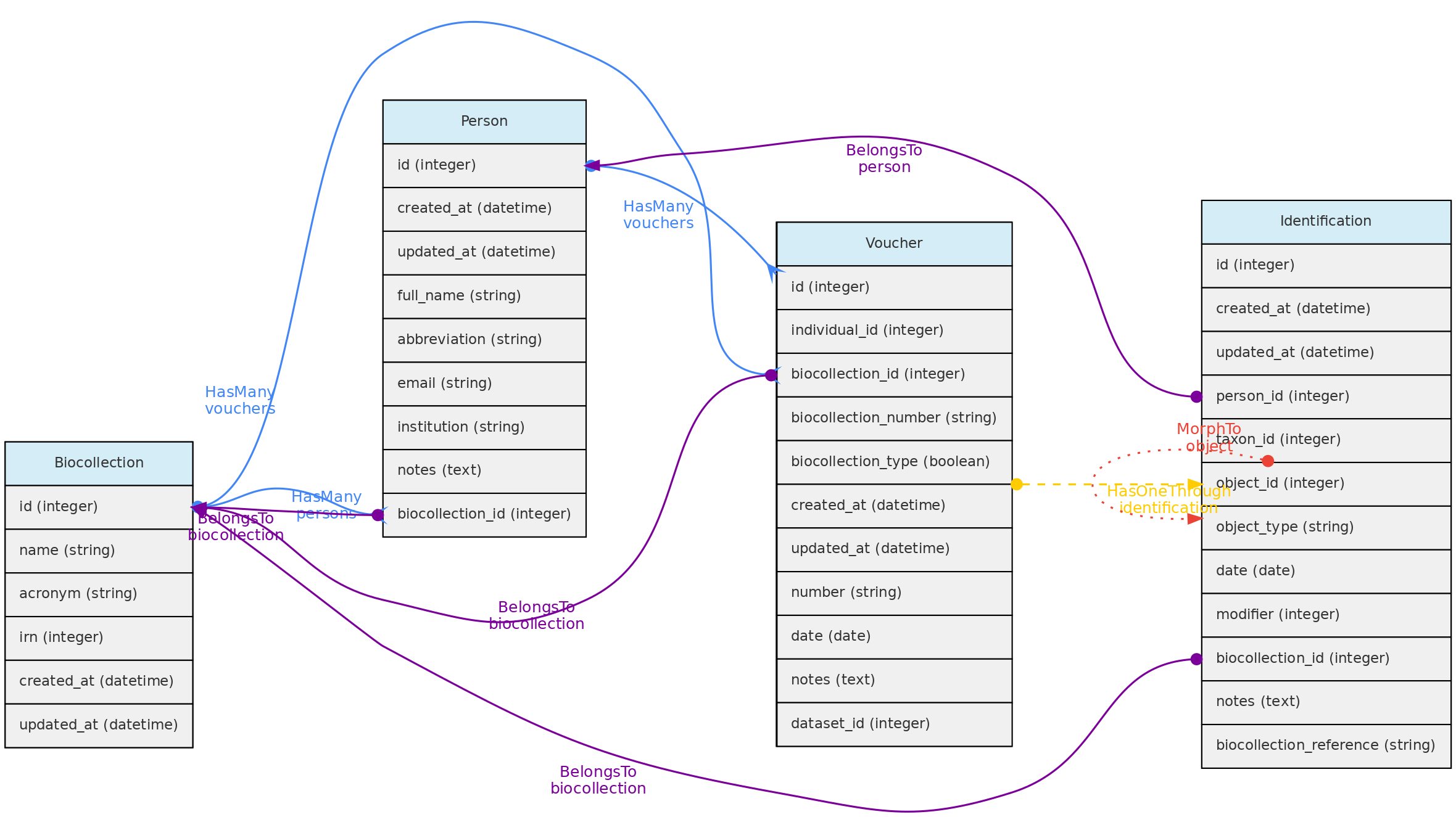
Data access - Full users can register BioCollections, but only super administrator can make a BioCollection manageable by the system. Removing BioCollections can be done if there are no Vouchers attached and if it is not administered by the system. If manageable, the model interacts with Users, which can be administrators (curators, anything can) or collaborators (can enter and edit data, but cannot delete records). Data from other Datasets can be part of the BioCollection, allowing users to have their complete data, but editing control of records is by the BioCollection authorized users.
Datasets
DataSets are groups of Measurements, Individuals, Vouchers and/or Media Files, and may have one or more Users administrators, collaborators or viewers. Administrators may set the privacy level to public access, restricted to registered users or restricted to authorized users or restricted to project users. This control access to the data within a dataset as exemplified in diagram below:
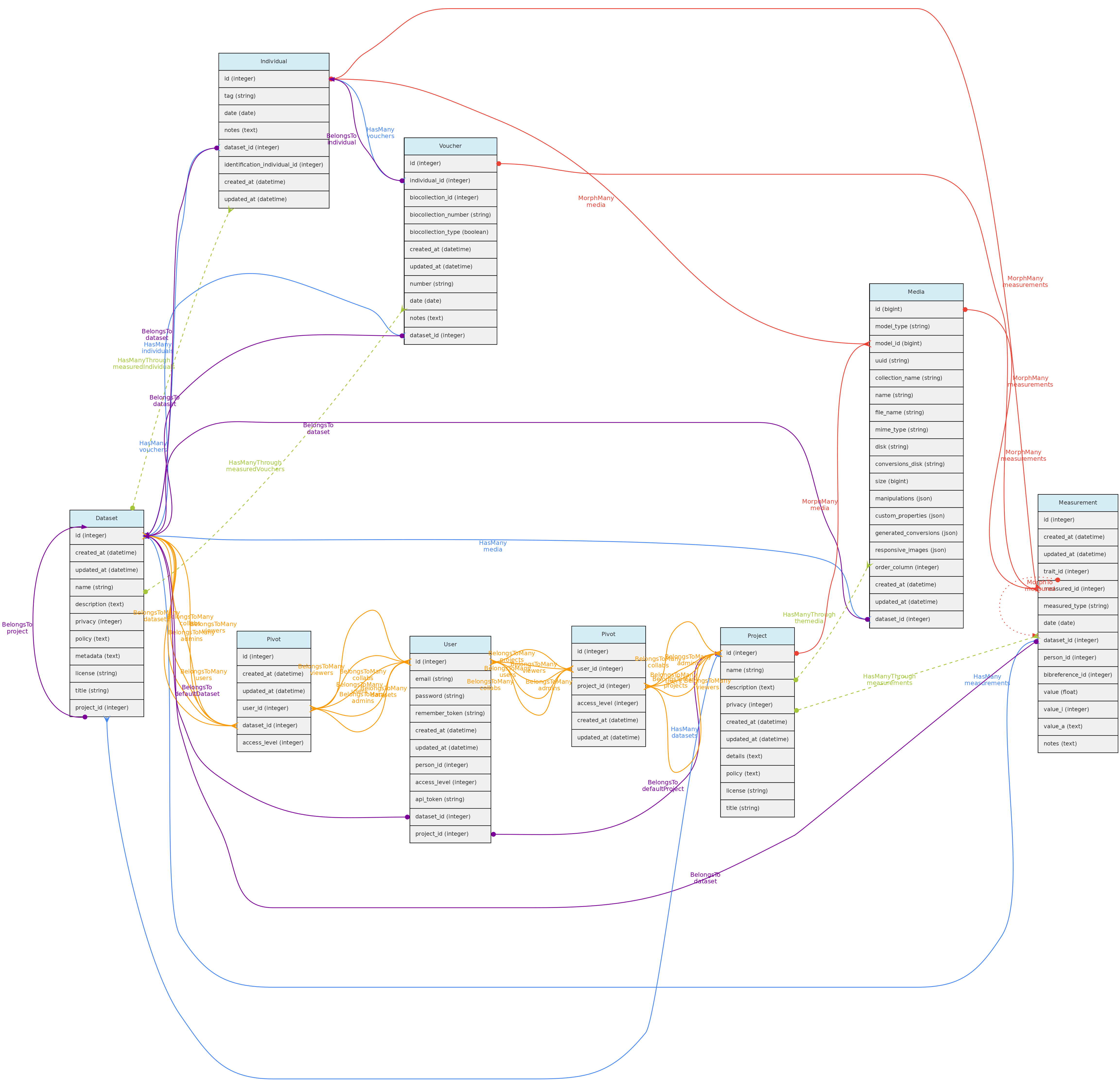
Datasets may also have many Bibliographic References, which together with fields policy, metadata permits to annotate the dataset with relevant information for data sharing:
* Link any publication that have used the dataset and optionally indicate that they are of mandatory citation when using the data;
* Define a specific data policy when using the data in addition to the a CreativeCommons.org public license;
* Detail any relevant metadata in addition to those that are automatically retrieved from the database like the definitions of the Traits measured.
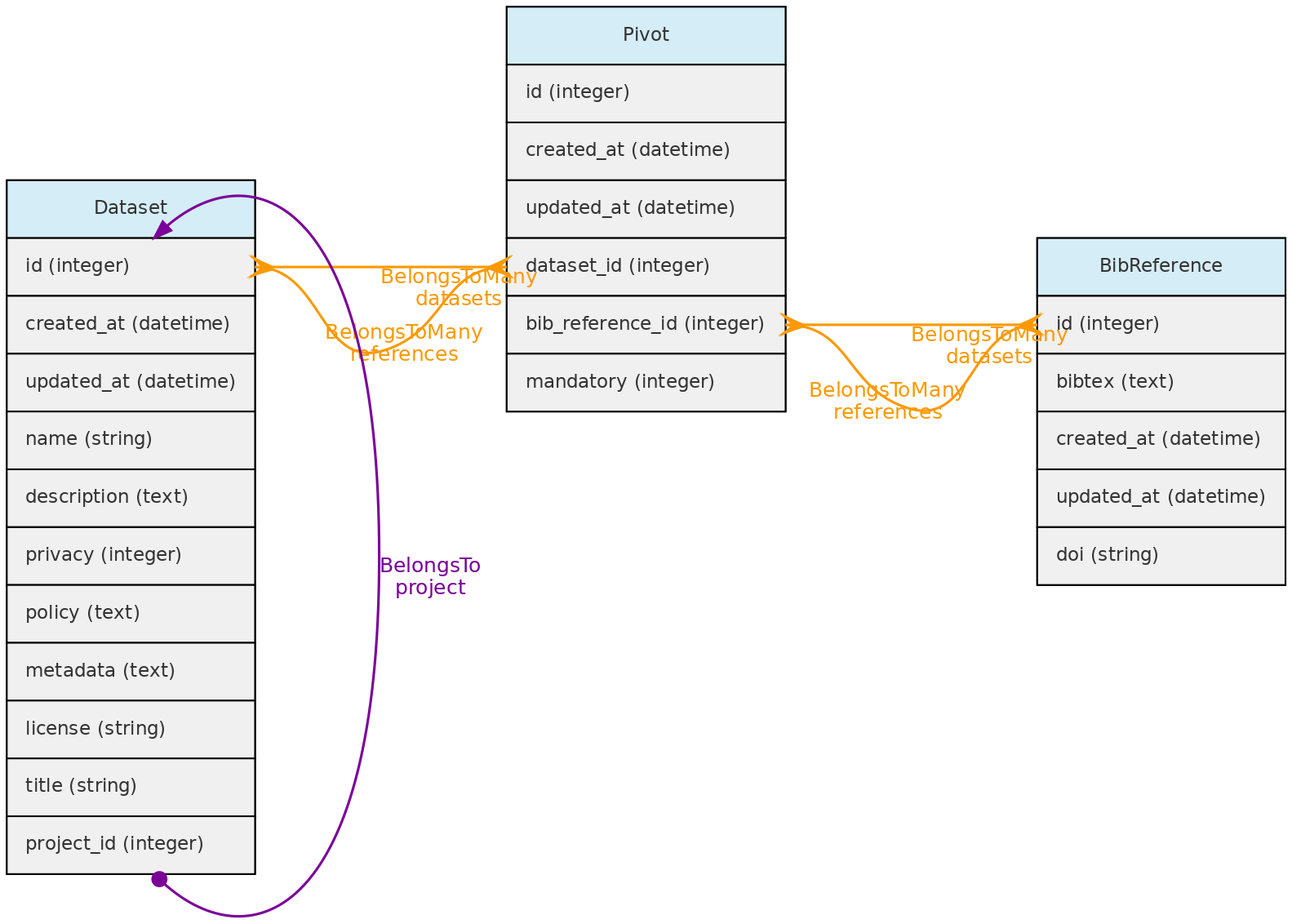
Projects
Projects are just groups of Datasets and interacts with Users, having administrators, collaborators or viewers. These users may control all datasets within the Project having a restricted to project users access policy.
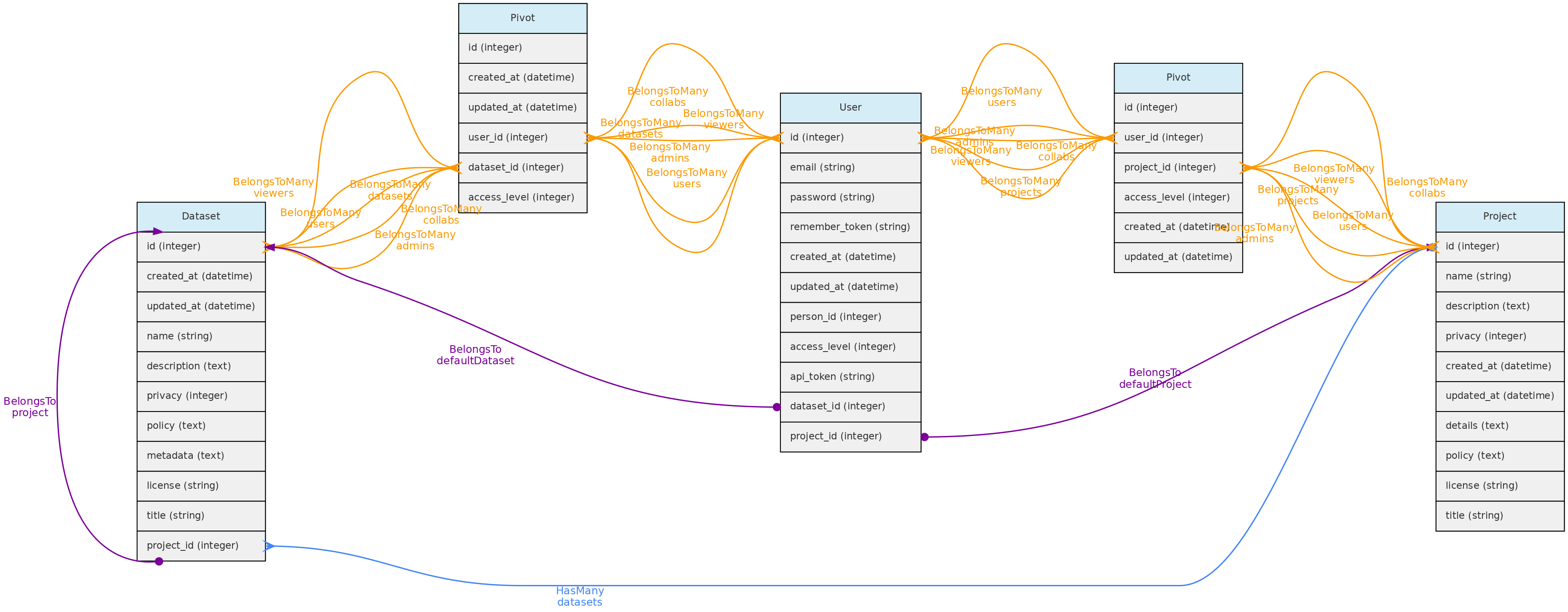
Users
The Users table stores information about the database users and administrators. Each User may be associated with a default Person. When this user enters new data, this person is used as the default person in forms. The person can only be associated to a single user.
There are three possible access levels for a user:
* Registered User (the lowest level) - have very few permissions
* Full User - may be assigned as administrators or collaborators to Projects and Datasets;
* SuperAdmin (the highest level). - superadmins have have access to all objects, regardless of project or dataset configuration and is the system administrator.
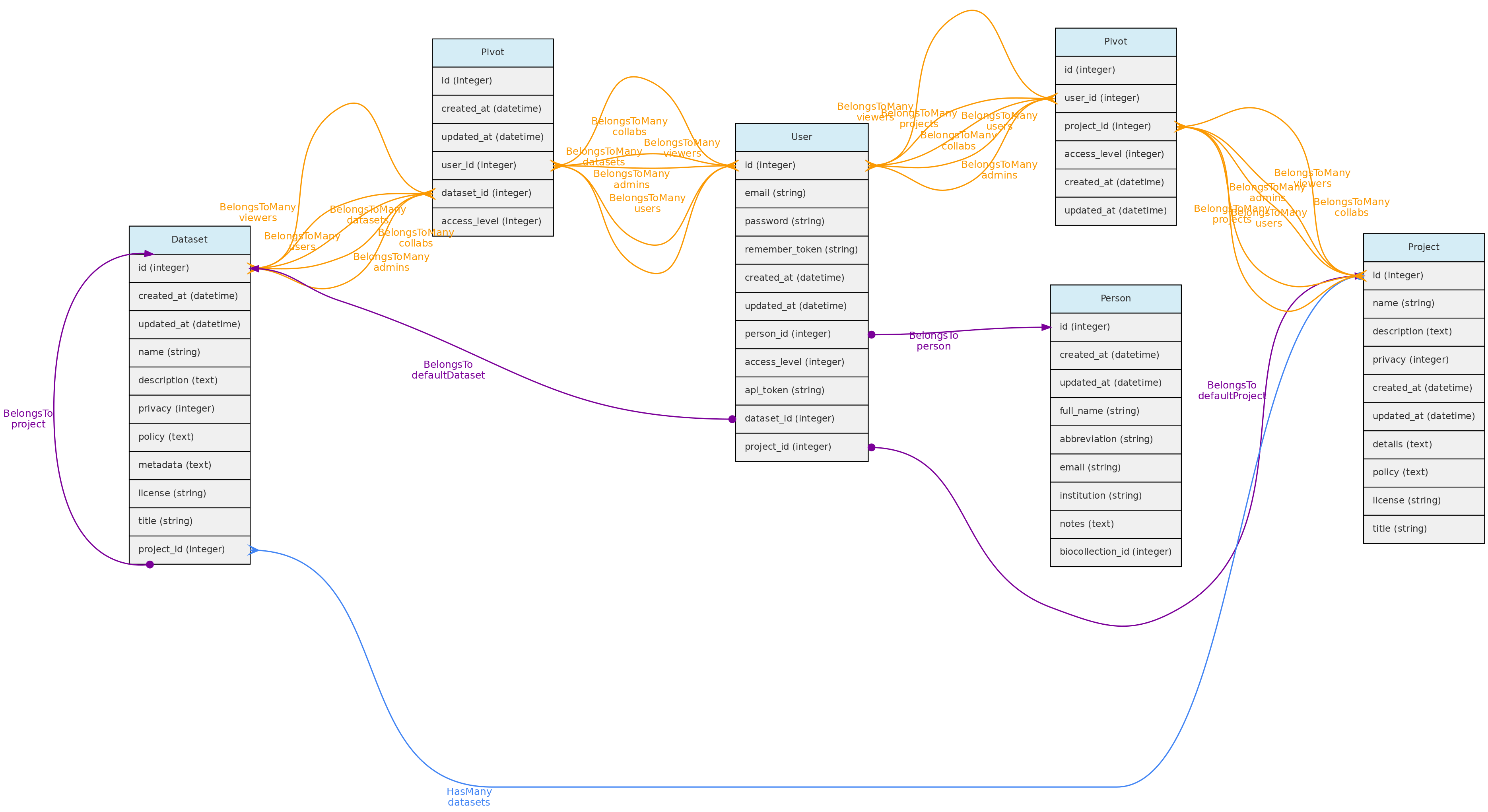
Each user is assigned to the registered user level when she or he registers in an OpenDataBio system. After that, a SuperAdmin may promote her/him to Full User or SuperAdmin. SuperAdmins also have the ability to edit other users and remove them from the database.
Every registered user is created along with a restricted Project and Dataset, which are referred to as her user Workspace. This allows users to import individual and voucher data before incorporating them into a larger project. [TO IMPLEMENT: export batches of objects from one project to another].
Data Access:users are created upon registration. Only administrators can update and delete user records.
User Jobs
The UserJob table is used to store temporarily background tasks, such as importing and exporting data. Any user is allowed to create a job; cancel their own jobs; list jobs that have not been deleted. The Job table contains the data used by the Laravel framework to interact with the Queue. The data from this table is deleted when the job runs successfully. The UserJob entity is used to keep this information, along with allowing for job logs, retrying failed jobs and canceling jobs that have not yet finished.
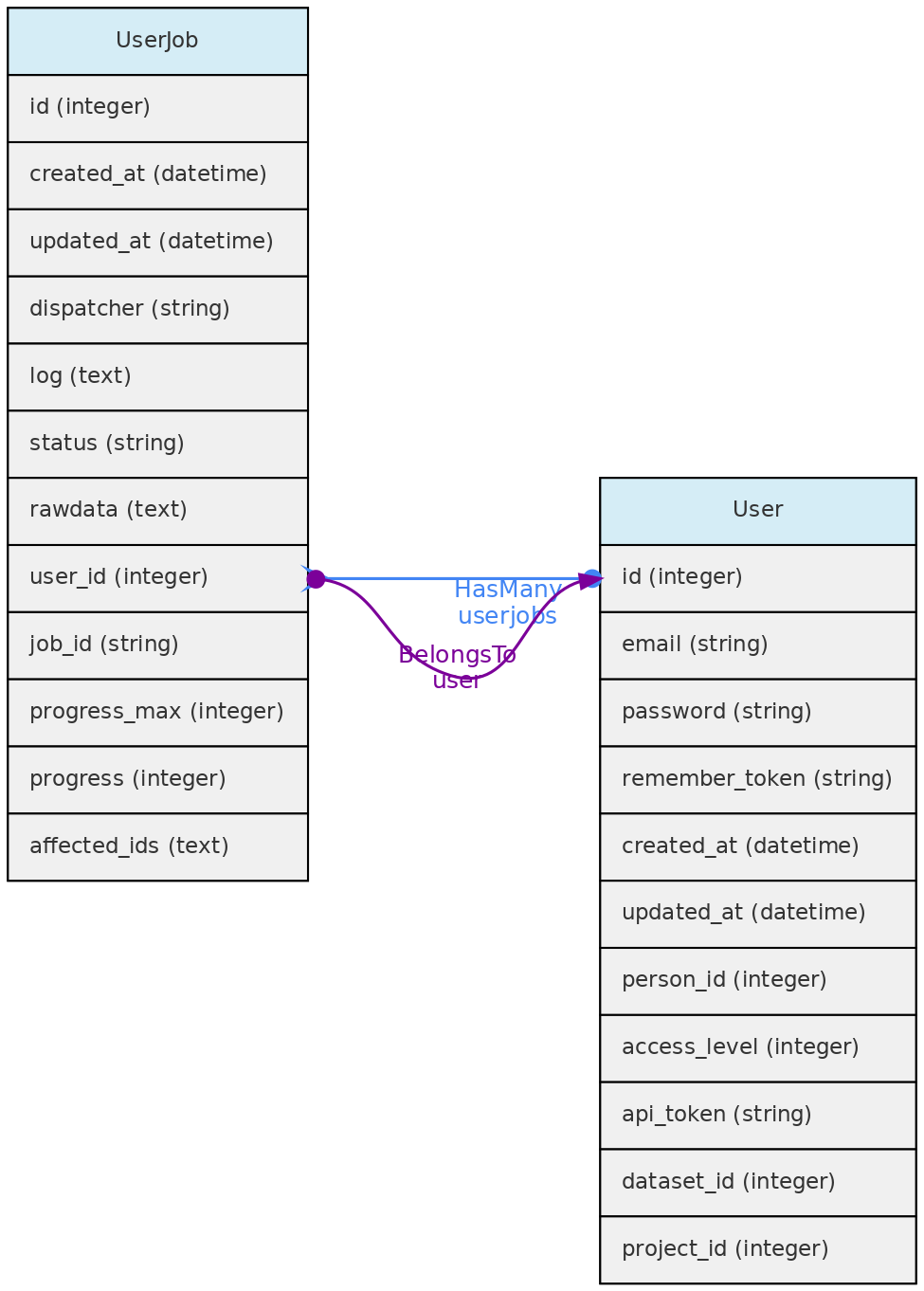
Data Access: Each registered user can see, edit and remove their own UserJobs.
4.4 - Auxiliary Objects
BibReference Model
The BibReference table contains basically BibTex formatted references stored in the bibtex column. You may easily import references into OpenDataBio by just specifying the doi, or simply uploading a bibtex record. These bibliographic references may be used to:
- Store references for Datasets - with the option of defining references for which citation is mandatory when using the dataset in publications; but all references that have used the dataset may be linked to the dataset; links are done with a Pivot table named
dataset_bibreference; - Store the references for Taxons:
- to specify the reference in which the Taxon name was described, currently mandatory in some Taxonomic journals like PhytoTaxa. This description reference is stored in the
bibreference_idof the Taxons table. - to register any reference to a Taxon name, which are then linked through a pivot table named
taxons_bibreference.
- to specify the reference in which the Taxon name was described, currently mandatory in some Taxonomic journals like PhytoTaxa. This description reference is stored in the
- Link a Measurement to a published source;
- Indicate the source of a Trait definition.
- Indicate mandatory citations for a Dataset, or link references using the data to a Dataset
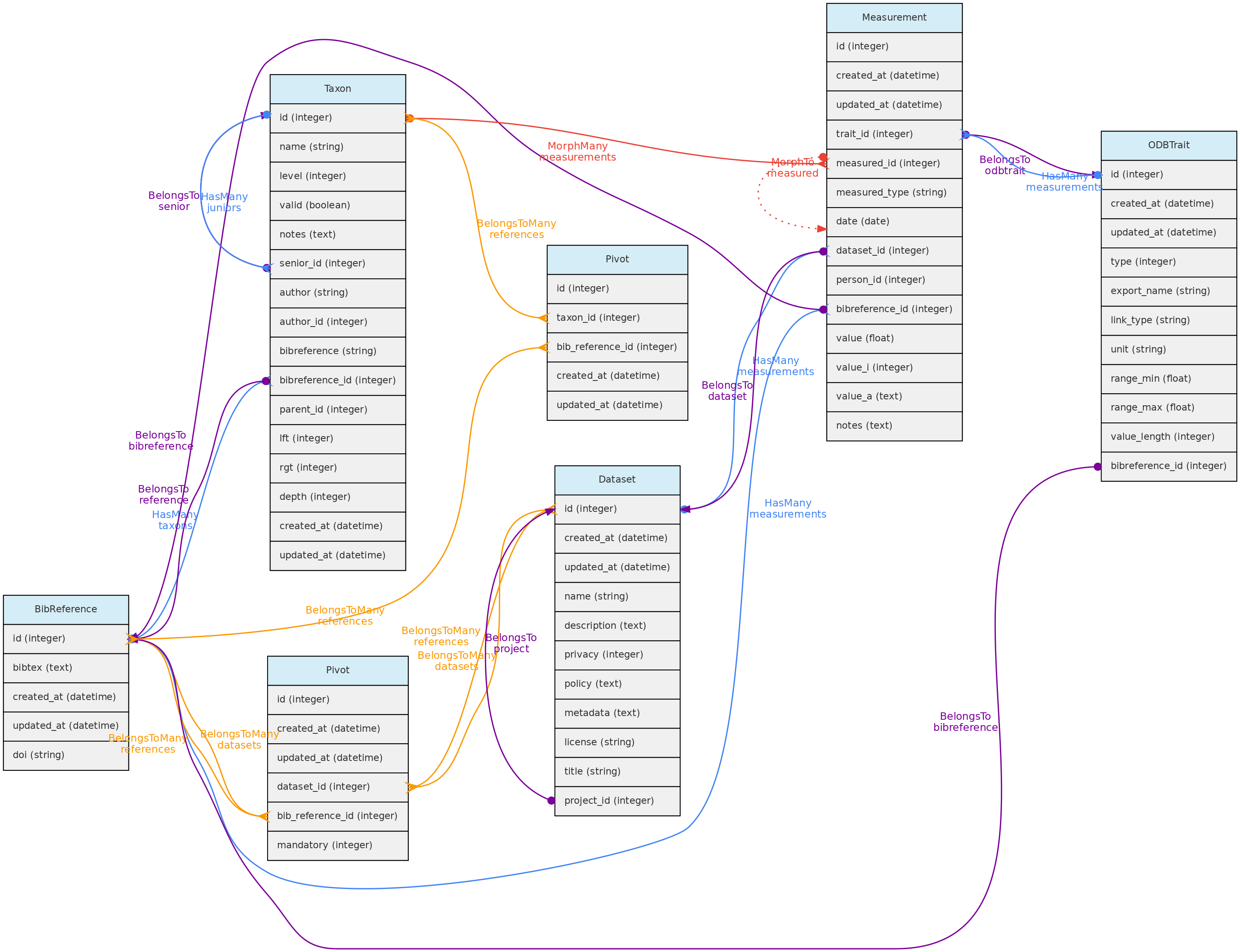 BibReference model and its relationships. Lines linking tables indicate the
BibReference model and its relationships. Lines linking tables indicate the methods implemented, with colors indicating different Eloquent relationships.
Bibreferences table
- The BibtexKey, authors and other relevant fields are extracted from the
bibtexcolumn. - The Bibtexkey must be unique in the database, and a helper function is be provided to standardize it with format
<von? last name> <year> <first word of title>. The “von part” of the name is the “von”, “di”, “de la”, which are part of the last name for some authors. The first word of the title ignores common stop-words such as “a”, “the”, or “in”. - DOIs for a BibReference may be specified either in the relevant BibTex field or in a separate text input, and are stored in the
doifield when present. An external API finds the bibliographic record when a user informs thedoi.
**Data access** [full users](/en/docs/concepts/data-access/#user) may register new references, edit references details and remove reference records that have no associated data. BibReferences have public access!
Identification Model
The Identification table represents the taxonomic identification of Individuals.
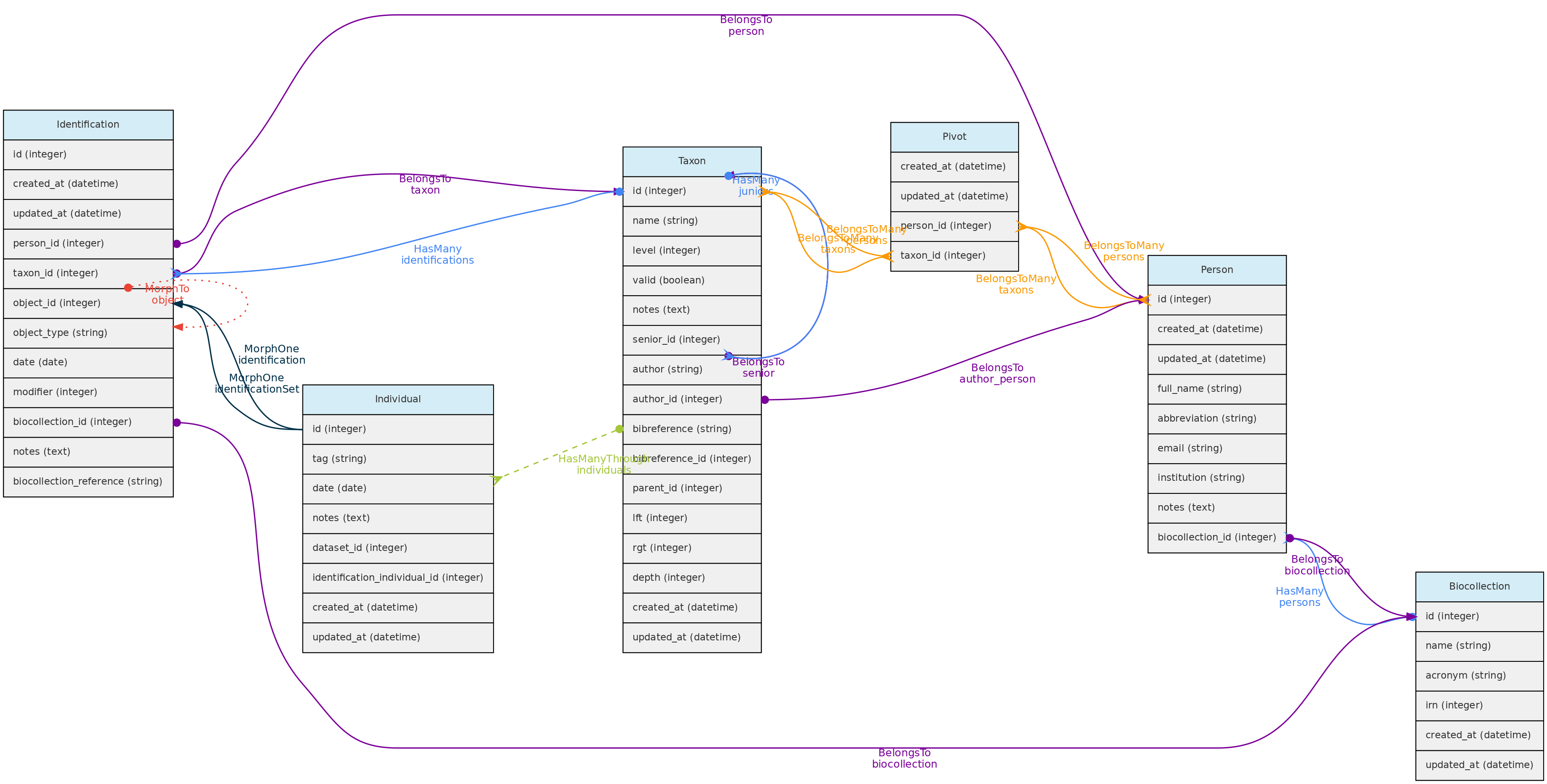 Identification model and its relationships. Lines linking tables indicate the
Identification model and its relationships. Lines linking tables indicate the methods implemented, with colors indicating different Laravel Eloquent relationships
Identifications table
- The Identification model includes several optional fields, but in addition to
taxon_id,person_id, the Person responsible for the identification, and the identificationdateare mandatory.
- The
datevalue may be an Incomplete Date, e.g. only the year or year+month may be recorded.
- The following fields are optional:
modifier- is a numeric code appending a taxonomic modifier to the name. Possible values ’s.s.’=1, ’s.l.’=2, ‘cf.’=3, ‘aff.’=4, ‘vel aff.’=5, defaults to 0 (none).notes- a text of choice, useful for adding comments to the identification.biocollection_idandbiocollection_reference- these fields are to be used to indicate that the identification is based upon comparison to a voucher deposited in a Biological Collection and creates a link between the Individual identified and the BioCollection specimen from which the identification is based upon.biocollection_idstores the Biocollection id, andbiocollection_referencethe unique identifier of the specimen compared, i.e. would be the equivalent of thebiocollection_numberof the Voucher model, but this reference does not need to be from a voucher registered in the database.
- The relationship with the Individual model is defined by a polymorphic relationship using fields
object_typeandobject_id[This could be replaced by an ‘individual_id’ in the identification table. The polymorphic relation inherited from a previous development version, kept because the Identification model may be used in the future to link Identifications to Measurements]. - Changes in identifications are audited for tracking change history
Data access: identifications are attributes of Individuals and do not have independent access!
Person Model
The Person object stores persons names, which may or may not be a User directly involved with the database. It is used to store information about people that are:
* collectors of Vouchers, Individuals and MediaFiles
* taxonomic determinators or identifiers of individuals;
* measurer of Measurements;
* authors for unpublished Taxon names;
* taxonomic specialists - linked with Taxon model by a pivot table named person_taxon;
* dataset authors - defining authors for the dynamic publication of datasets;
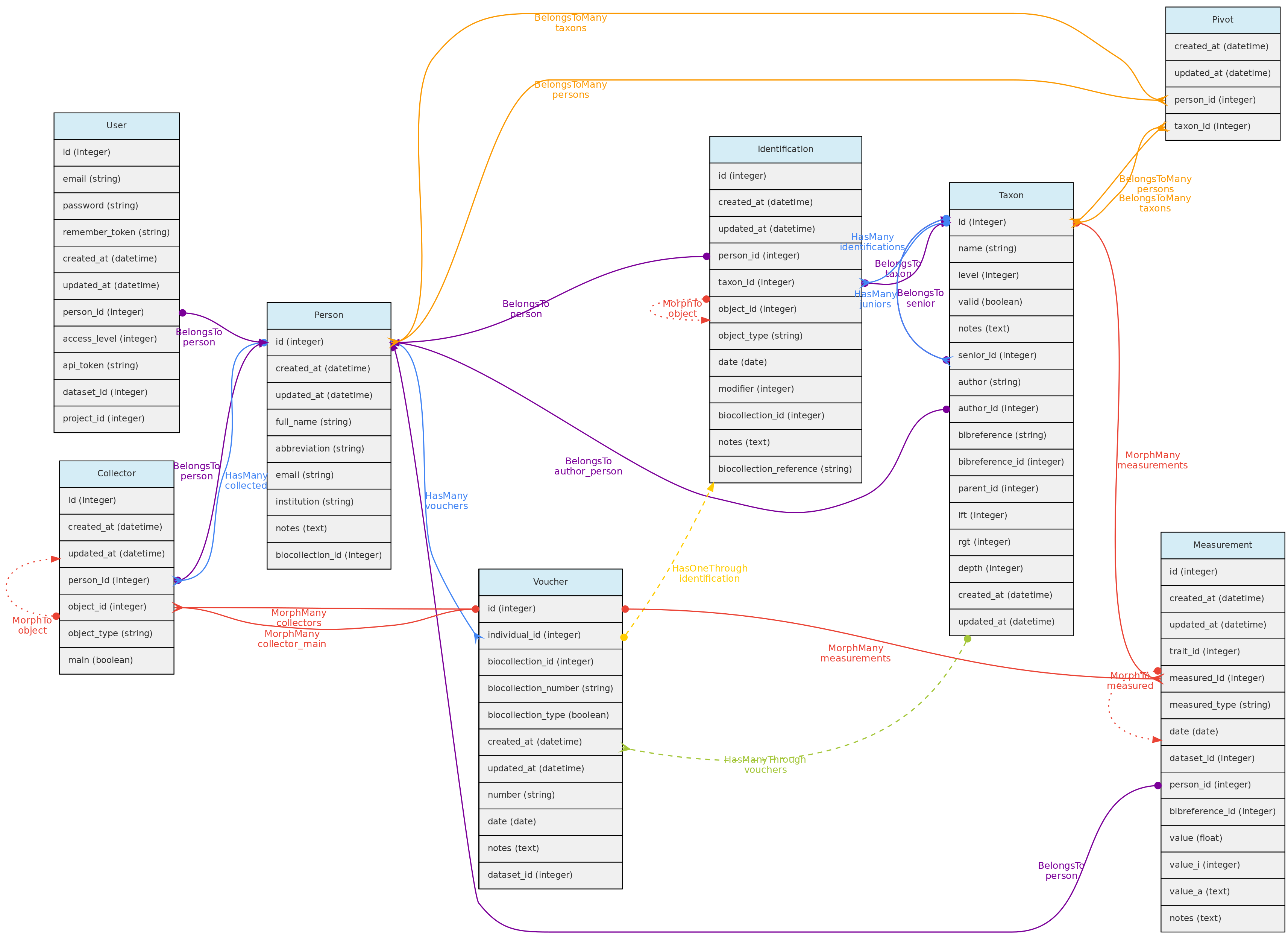 Person model and its relationships. Lines linking tables indicate the
Person model and its relationships. Lines linking tables indicate the methods implemented, with colors indicating different types of Laravel Eloquent methods, solid lines the direct and dashed the indirect relationships
Persons table
- mandatory columns are the person
full_nameandabbreviation; - when registering a new person, the system suggests the name
abbreviation, but the user is free to change it to better adapt it to the usual abbreviation used by each person. The abbreviation must be unique in the database, duplicates are not allowed in the Persons table. Therefore, two persons with the exact same name must be differentiated somehow in theabbreviationcolumn. - The
biocollection_idcolumn of the Persons table is used to list to which Biocollection a person is associated, which may be used when the Person is also a taxonomic specialist. - Additionally, the
emailandinstitutionthe person belongs to may also be informed. - Each user can be linked to a Person by the
person_idin the User table. This person is then used the ‘default’ person when the user is logged into the system.
**Data access** [full users](/en/docs/concepts/data-access/#user) may register new persons and edit the persons they have inserted and remove persons that have no associated data. Admins may edit any Person. Persons list have public access.
Media Model
Media files are similar to measurements in that they might be associated with any core object. Media files may be images (jpeg, png, gif, tif), video or audio files and can be made freely accessible or placed in a Dataset with a defined access policy. A CreativeCommons.org license must be assigned to them. Media files may be tagged, i.e. you may define keywords to them, allowing to query them by Tags. For example, an individual image may be tagged with ‘flowers’ or ‘fruits’ to indicate what is in the image, or a tag that informs about image quality.
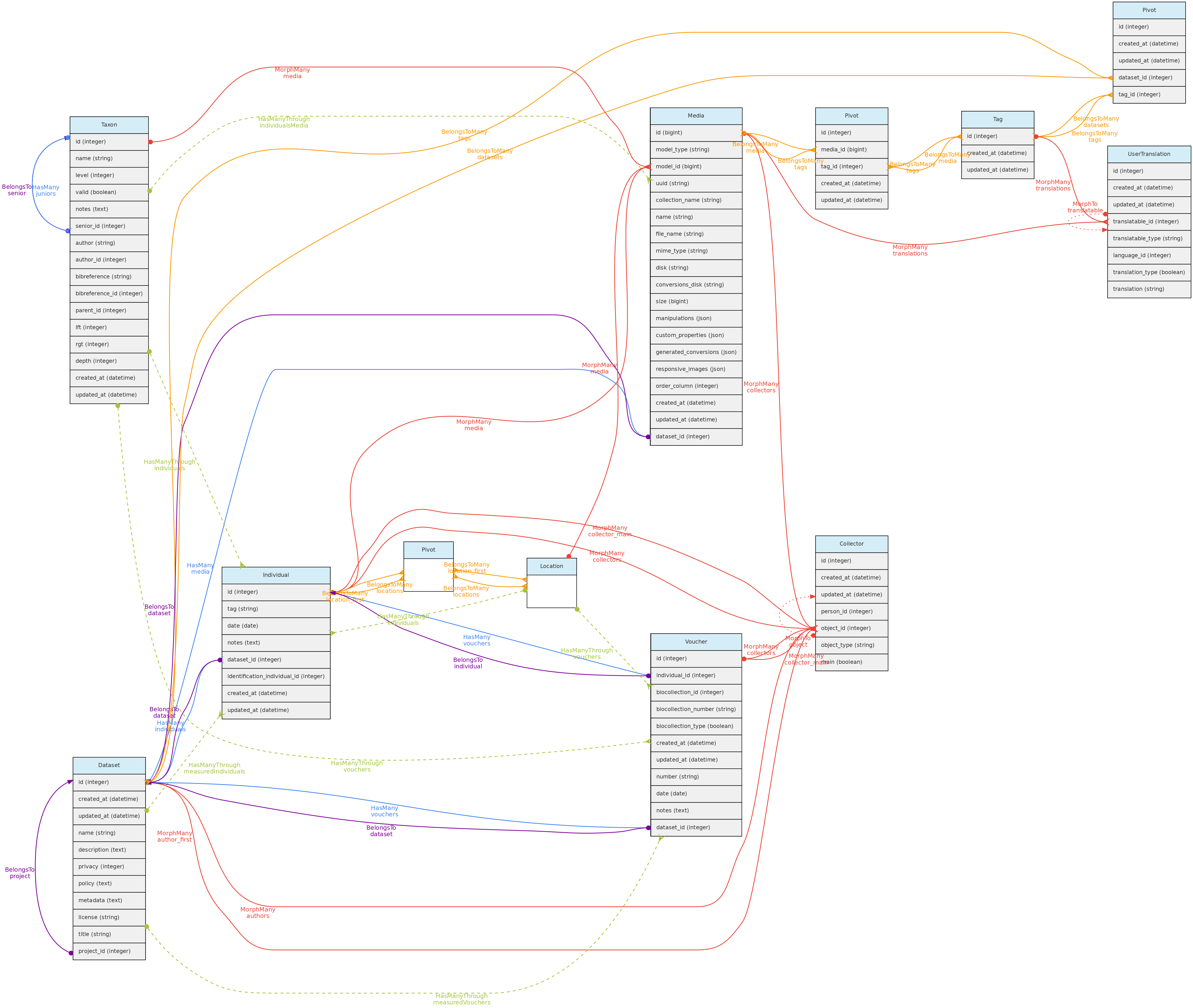
- Media files (image, video, audio) are linked to the Core-Objects through a polymorphic relationship defined by columns
model_idandmodel_type. - Multiple Persons may be associated with the Media for credits, these are linked with the Collectors table and its polymorphic relationship structure.
- A Media may have a
descriptionin each language configured in the Language table, which will be stored in theuser_translationstable, which relates to the Tag model through a polymorphic relationship. Inputs for each language are shown in the web-interface forms. - Media files are not stored in the database, but in the server storage folder.
- It is possible to batch upload media files through the web interface, requiring also a file informing the objects to link the media with.
Data access full users may register media files and delete the ones they have inserted. If Media is in a Dataset, dataset admins may delete the media in addition to the user. Media files have public access, except when linked to a Dataset with access restrictions.
Tag Model
The Tag model allows users to define translatable keywords that may be used to flag Datasets, Projects or MediaFiles. The Tag model is linked with these objects through a pivot table for each, named dataset_tag, project_tag and media_tag, respectively.
A Tag may have name and description in each language configured in the Language table, which will be stored in the user_translations table, which relates to the Tag model through a polymorphic relationship. Inputs for each language are shown in the web-interface forms.
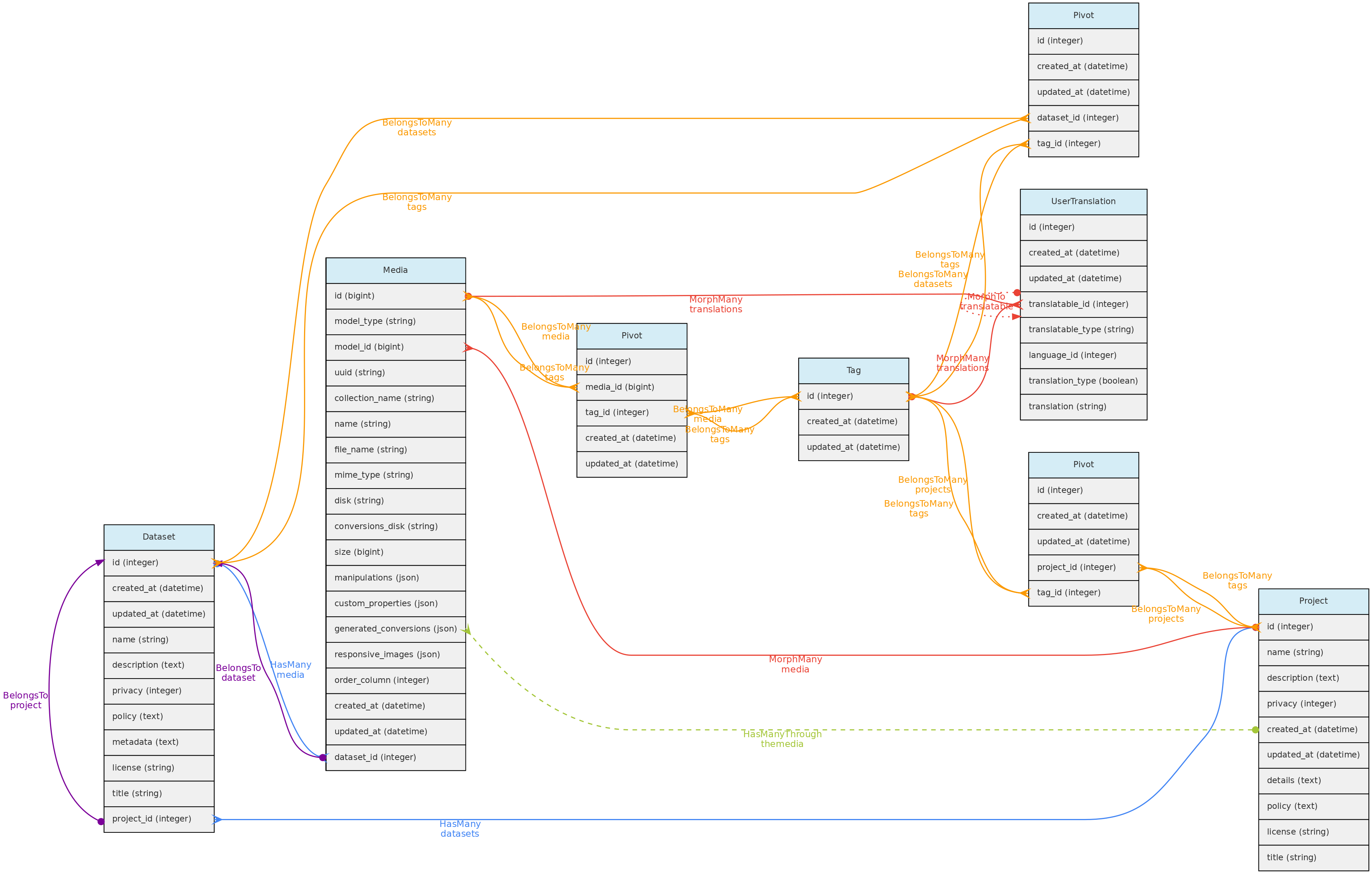
Data access full users may register tags, edit those they have inserted and delete those that have not been used. Tags have public access as they are just keywords to facilitate navigation.
User Translation Model
The UserTranslation model translates user data: Trait and Trait Categories names and descriptions, MediaFiles descriptions and Tags. The relations between these models are established by polymorphic relations using fields translatable_type and translatable_id. This model permits translations to any language listed in the Language table, which is currently only accessible for insertion and edition directly in the SQL database. Input forms in the web interface will be listed for registered Languages.
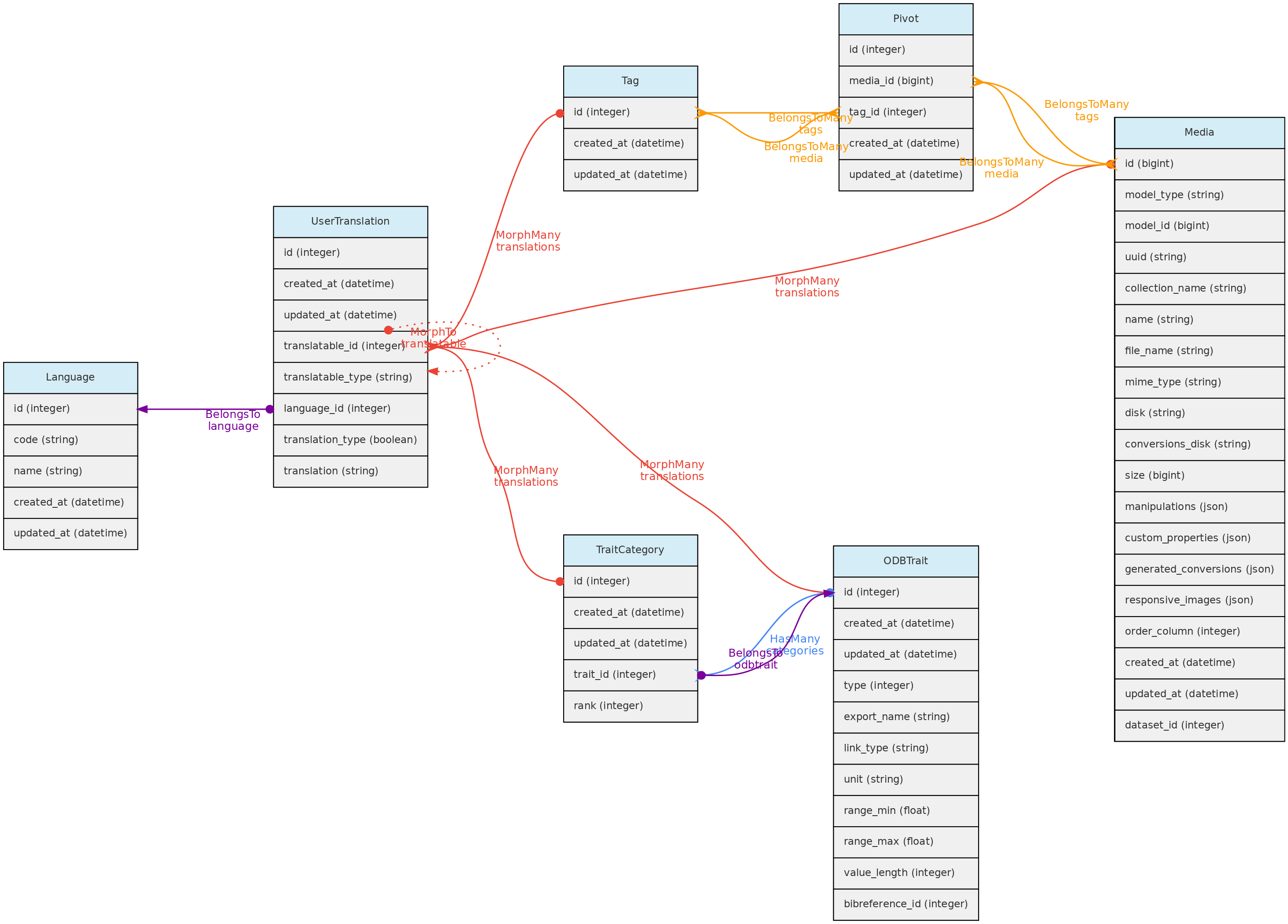
Vernacular - Popular Name
The Vernacular and VernacularCitation templates allow you to record popular names of organisms by relating them to Taxons and/or Individuals. Names must be unique in the Vernacular table, and each record can be linked to multiple Taxa and/or Individuals, depending on the information sources. Each record can also have one or more citations (citation text + BibReference + note).
Incomplete Dates
Dates for Vouchers, Individuals, Measurements and Identifications may be Incomplete, but at least year is mandatory in all cases. The date columns in the tables are of ‘date’ type, and incomplete dates are stored having 00 in the missing part: ‘2005-00-00’ when only year is known; ‘1988-08-00’ when only month is known.
Auditing
Modifications in database records are logged to the activity_log table. This table is generated by the package ActivityLog. The activities are shown in a ‘History’ link provided in the Show.view of the models.
- The package stores changes as json in the
propertiesfield, which contains two elements:attributesandold, which are basically the new vs old values that have been changed. This structure must be respected. - Class ActivityFunctions contain custom functions to read the the properties Json record stored in the
activity_logtable and finds the values to show in the History datatable; - Most changes are logged by the package as a ’trait’ called within the Class. These allow to automatically log most updates and are all configured to log only the fields that have changed, not entire records (option
dirty). Also record creation are not logged as activity, only changes. - Some changes, like Individual and Vouchers collectors and identifications are manually logged, as they involve related tables and logging is specified in the Controller files;
- Logging contain a
log_namefield that groups log types, and is used to distinguish types of activity and useful to search the History Datatable; - Two special logging are also done:
- Any Dataset download is logged, so administrators may track who and when the dataset was downloaded;
- Any Dataset request is also logged for the same reason
The clean-command of the package SHOULD NOT be used during production, otherwise will just erase all logged changes. If run, will erase the logs older than the time specified in the /config/activitylog.php file.
The ActivityLog table has the following structure:
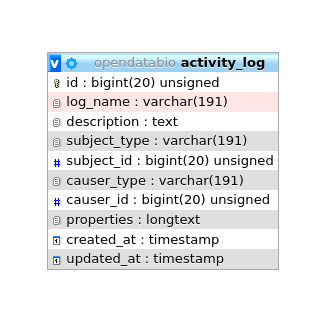
causer_typeandcauser_idwill be the User that made the changesubject_typeandsubject_idwill the model and record changedlog_name- to group logs together and permits queriesdescription- somewhat redundant with log_name in a OpenDataBio context.properties- stores the changes, for example, and identification change will have a log like:
{
"attributes":
{
"person_id":"2",
"taxon_id":"1424",
"modifier":"2",
"biocollection_id":"1",
"biocollection_reference":"1234",
"notes":"A new fake note has been inserted",
"date":"2020-02-08"},
"old":{
"person_id":674,
"taxon_id":1413,
"date":"1995-00-00",
"modifier":0,
"biocollection_id":null,
"notes":null,
"biocollection_reference":null
}
}
5 - Contribution Guidelines
Report bugs & suggest improvements
Post an issue on one of the GitLab repositories below, depending on the issue.
Before posting, check if an open issue does already contains what you want to report, ask or propose.
Tag your issue with one or more appropriate labels.
Issues for the main repository
Issues for the R package
Issues for this documentation site
Collaborate with development, language translations and docs
We expect this project to grow collaboratively, required for its development and use in the long term. Therefore, developer collaborators are welcome to help fixing and improving OpenDataBio. The issues list is a place to start to know what is needed.
The following guidelines are recommend if you want to collaborate:
- Communicate with the OpenDataBio repository maintainer indicating which issues you want to work on and join the development team.
Forkthe repository- Create a
branchto commit your modifications or additions - When happy with results, make a pull request to ask the project maintainer to review your contribution and merge it to the repository. Consult GitLab Help for more information on using pull requests.
Programming directives
- Use the docker installation for development, which shared among all developers facilitates debug. The Laravel-Datatables library is incompatible with
php artisan serve, so this command should not be used. - This software should adhere to Semantic Versioning, starting from version 0.1.0-alpha1. The companion R package and the Documentation (this site) should follow a similar versioning scheme. When changing version, a release tag must be created with the old version.
- All variables and functions should be named in English, with entities and fields related to the database being named in the singular form. All tables (where appropriate) should have an “id” column, and foreign keys should reference the base table with “_id” suffix, except in cases of self-joins (such as “taxon.parent_id”) or polymorphic foreign keys. The id of each table has type INT and should be autoincrementing.
- Use laravel migration class to add any modification to the database structure. Migration should include, if apply, management of existing data.
- Use camelCase for methods (i.e. relationships) and snake_case for functions.
- Document the code with comments and create documentation pages if necessary.
- There should be a structure to store which Plugins are installed on a given database and which are the compatible system versions.
- This system uses Laravel Mix to compile the SASS and JavaScript code used. If you add or modify these
npm run prodafter making any change to these files.
Collaborate with the docs
We welcome Tutorials for dealing with specific tasks.
To create a tutorial:
Forkthe documentation repository. When cloning this repository or a fork, include the submodule option to also get the included Docsy theme repository. You will need Hugo to run this site in your localhost.- Create a
branchto commit your modifications or additions - Add your tutorial:
- Create a folder within the
contents/{lang}/docs/Tutorialsusing kebab-case for the folder name. Ex.first-tutorial - You may create a tutorial in single language, or on multiple languages. Just place it in the correct folder
- Within the created folder, create a file named
_index.mdand create the markdown content with your tutorial. - You may start copying the content of an existing tutorial.
- When happy with results, make a pull request to ask the project maintainer to review your contribution and merge it to the repository. Consult GitLab Help for more information on using pull requests.
Collaborate with translations
You may help with translations for the web-interface or the documentation site. If want to have a new language for your installation, share your translation, creating a pull request with the new files.
New language for the web-interface:
- fork and create a branch for the main repository
- create a folder for the new language using the ISO 639-1 Code within the
resources/langfoldercd opendatabio cd resources/lang cp en es - translate all the values for all the variables within all the files in the new folder (may use google translation to start, just make sure variable names are not translated, otherwise, it will not work)
- add language to array in
config/languages.php - add language to database language table creating a laravel migration
- make a pull request
New language for the documentation site
- fork and create a branch for the documentation repository
- create a folder for the new language using the ISO 639-1 Code within the
contentfoldercd opendatabio.gitlab.io cd contents cp pt es - check all files within the folder and translate where needed (may use google translation to start, just make sure to translate only what can be translated)
- push to your branch and make a pull request to the main repository
Polymorphic relations
Some of the foreign relations within OpenDataBio are mapped using Polymorphic relations. These are indicated in a model by having a field ending in _id and a field ending in _type. For instance, all Core-Objects may have Measurements, and these relationships are established in the Measurements table by the measured_id and the measured_type columns, the first storing the related model unique id, the second the measured model class in strings like ‘App\Models\Individual’, ‘App\Models\Voucher’, ‘App\Models\Taxon’, ‘App\Models\Location’.
Data model images
Most figures for explaining the data model were generated using Laravel ER Diagram Generator, which allows to show all the methods implemented in each Class or Model and not only the table direct links:
To generate these figures a custom php artisan command was generated. These command is defined in file app/Console/Commands/GenerateOdbErds.php.
To update the figures follow the following steps:
- Figures are configure in the
config/erd-generator-odb.phpfile. There are many additional options for customizing the figures by changing or adding graphviz variables to theconfig/erd-generator-base.phpfile. - The custom command is
php artisan odb:erd {$model}, where model is the key of the arrays inconfig/erd-generator-odb.php, or the word “all”, to regenerate all doc figures.
cd opendatabio
make ssh
php artisan odb:erd all
- Figures will be saved in
storage/app/public/dev-imgs - Copy the new images to the documentation site. They need to be placed within
contents/{lang}/concepts/{subfolder}for all languages and in the respective sub-folders.
6 - Tutorials
Find here working examples of using OpenDataBio through the web-interface or through the OpenDataBio R package. See Contribution guidelines if you want to contribute with a tutorial.
6.1 - Getting data with OpenDataBio-R
The Opendatabio-R package was created to allow users to interact with an OpenDataBio server, to both obtain (GET) data or to import (POST) data into the database. This tutorial is a basic example of how to get data.
Set up the connection
- Set up the connection to the OpenDataBio server using the
odb_config()function. The most important parameters for this function arebase_url, which should point to the API url for your OpenDataBio server, andtoken, which is the access token used to authenticate your user. - The
tokenis only need to get data from datasets that have one of the restricted access policies. Data from datasets of public access can be extracted without the token specification. - Your token is avaliable in your profile in the web interface
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ"
cfg = odb_config(base_url=base_url, token = token)
More advanced configuration involves setting a specific API version, a custom User Agent, or other HTTP headers, but this is not covered here.
Test your connection
The function odb_test() may be used to check if the connection was successful, and whether
your user was correctly identified:
odb_test(cfg)
#will output
Host: https://opendb.inpa.gov.br/api/v0
Versions: server 0.9.1-alpha1 api v0
$message
[1] "Success!"
$user
[1] "admin@example.org"
As an alternative, you can specify these parameters as systems variables. Before starting R, set this up on your shell (or add this to the end of your .bashrc file):
export ODB_TOKEN="YourToken"
export ODB_BASE_URL="https://opendb.inpa.gov.br/api"
export ODB_API_VERSION="v0"
GET Data
See the GET API Quick Reference for a complete list of endpoints and request parameters. Also see the generic parameters, especially
save_jobwhich is important for downloading large datasets.
For publicly accessible data the token is optional. Below are some examples. Follow a similar reasoning to use the other endpoints. See the R package help for all available odb_get_{endpoint} functions.
Getting Taxon names
See GET API Taxon Endpoint request parameters and a list of response fields.
base_url="https://opendb.inpa.gov.br/api"
cfg = odb_config(base_url=base_url)
#get id for a taxon
mag.id = odb_get_taxons(params=list(name='Magnoliidae',fields='id,name'),odb_cfg = cfg)
#use this id to get all descendants of this taxon
odb_taxons = odb_get_taxons(params=list(root=mag.id$id,fields='id,scientificName,taxonRank,parent_id,parentName'),odb_cfg = cfg)
head(odb_taxons)
If the server used the seed data provided and the default language is portuguese, the result will be:
id scientificName taxonRank parent_id parentName
1 25 Magnoliidae Clado 20 Angiosperms
2 43 Canellales Ordem 25 Magnoliidae
3 62 Laurales Ordem 25 Magnoliidae
4 65 Magnoliales Ordem 25 Magnoliidae
5 74 Piperales Ordem 25 Magnoliidae
6 93 Chloranthales Ordem 25 Magnoliidae
Getting Locations
See GET API Location Endpoint request parameters and a list of response fields. See also the POST Locations-Validation Endpoint if you have latitude and longitude and wants to validade the geometries and find where the point falls.
Get some fields listing all Conservation Units (adm_level==99) registered in the server:
base_url="https://opendb.inpa.gov.br/api"
cfg = odb_config(base_url=base_url)
odblocais = odb_get_locations(params = list(fields='id,name,parent_id,parentName',adm_level=99),odb_cfg = cfg)
head(odblocais)
If the server used the seed data provided and the default language is portuguese, the result will be:
id name
1 5628 Estação Ecológica Mico-Leão-Preto
2 5698 Área de Relevante Interesse Ecológico Ilha do Ameixal
3 5700 Área de Relevante Interesse Ecológico da Mata de Santa Genebra
4 5703 Área de Relevante Interesse Ecológico Buriti de Vassununga
5 5707 Reserva Extrativista do Mandira
6 5728 Floresta Nacional de Ipanema
parent_id parentName
1 6 São Paulo
2 6 São Paulo
3 6 São Paulo
4 6 São Paulo
5 6 São Paulo
6 6 São Paulo
Locations as spatial objects in R
To obtain a spatial object in R, use the sf package. The example below plots a plot and its subplots and also exports the locations as both, kml and shapefile.
library(sf)
library(opendatabio)
#download all plot locations
cfg <- odb_config(base_url = "https://opendb.inpa.gov.br/api")
#get a large main plot
parcela = odb_get_locations(params = list(fields='all',name='Parcela 25ha'), odb_cfg = cfg)
parcela$type = 'main plot'
#get subplots
subplots = odb_get_locations(params = list(fields='all',location_root=parcela$id), odb_cfg = cfg)
subplots = subplots[subplots$adm_level==100 & subplots$id!=parcela$id,]
subplots$type = 'sub plot'
#convert footprintWKT to sf geometries
geoms <- st_as_sfc(parcela$footprintWKT, crs = 4326)
parcela_sf <- st_sf(parcela, geometry = geoms)
geoms <- st_as_sfc(subplots$footprintWKT, crs = 4326)
subplots_sf <- st_sf(subplots, geometry = geoms)
#print a figure
png("plots_with_subplots.png", width = 15, height = 15,units='cm',res=300)
par(mar=c(2,2,3,2))
plot(st_geometry(subplots_sf),border='green',main = parcela$locationName)
plot(st_geometry(parcela_sf),border='red',add=T)
labs = gsub("Quadrat ","",subplots_sf$locationName)
text(
st_coordinates(st_centroid(subplots_sf)),
labels = labs,
cex = 0.2, col = "blue"
)
dev.off()
#save as kml
locais = rbind(parcela_sf,subplots_sf)
locais$name <- locais$locationName
cols_to_include <- setdiff(names(locais), c("footprintWKT",'locationName'))
locais_kml <- locais[, c("name", cols_to_include[cols_to_include != "name"])]
st_write(locais_kml, "plots_and_subplots.kml", layer=parcela$locationName, driver = "KML", delete_dsn = TRUE)
#save as shapefile
st_write(locais_kml, "plots_and_subplots.shp", layer=parcela$locationName, delete_layer = TRUE)
Figure generated:
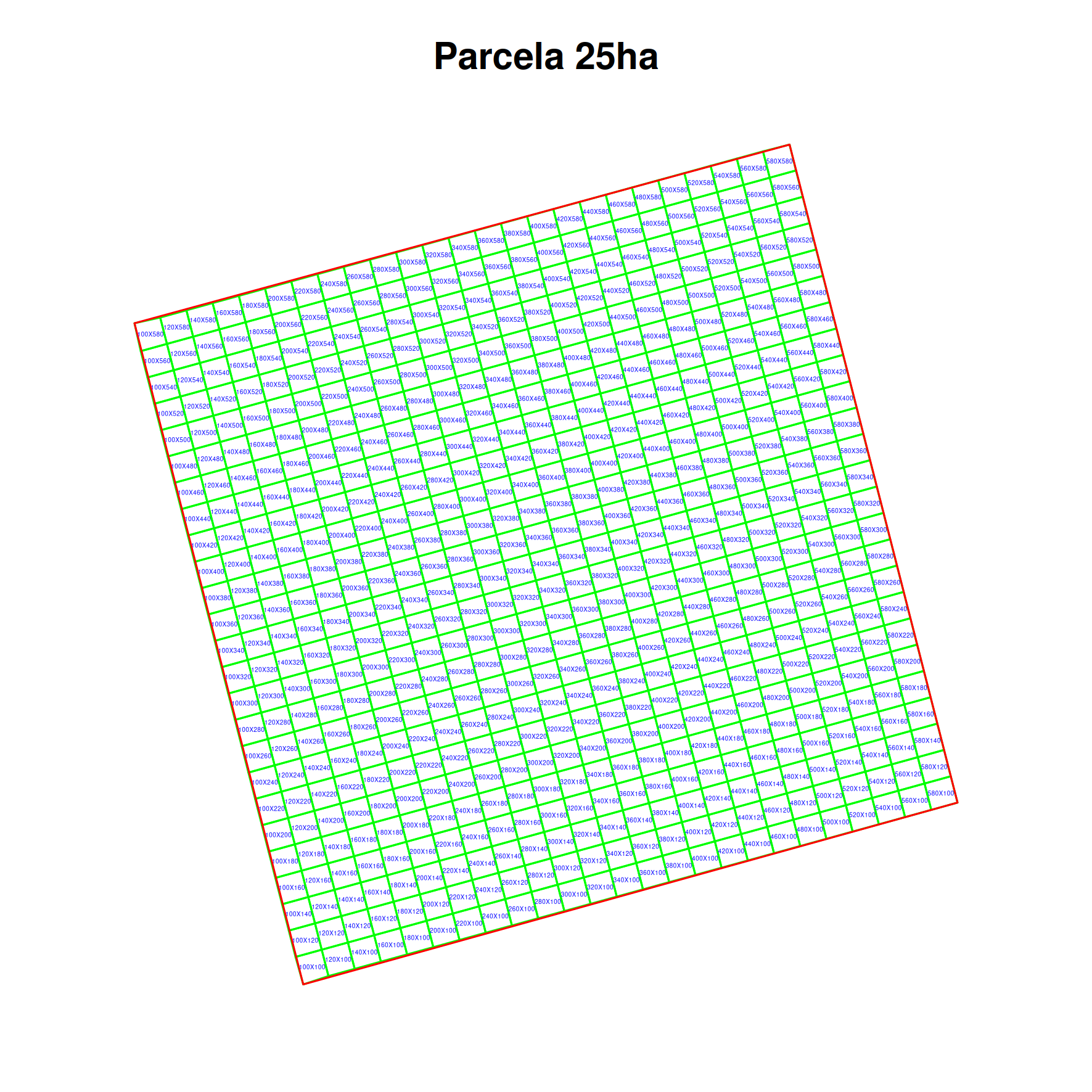

Validating point geometries
See the POST Locations-Validation Endpoint.
#conect to database
library(opendatabio)
base_url="http://localhost/opendatabio/api"
token ="your token is mandatory in this case"
cfg = odb_config(base_url=base_url, token = token)
odb_test(cfg)
#fake data
dados = data.frame(
latitude = sample(seq(-2,2,by=0.00001),10),
longitude = sample(seq(-60,-59,by=0.00001),10)
)
#submit job for validation
jb = odb_validate_locations(dados,odb_cfg = cfg)
#monitor job execution
odb_get_jobs(params=list(id=jb$id),odb_cfg = cfg)
#get results
dadosValidados = odb_get_jobs(params=list(id=jb$id,get_file=T),odb_cfg = cfg)
head(dados)
latitude longitude
1 0.12975 -59.65745
2 1.77469 -59.77757
3 -0.89154 -59.80179
4 -1.25632 -59.87084
5 0.77085 -59.22740
6 -0.74237 -59.64591
head(dadosValidados)
latitude longitude withinLocationName withinLocationParent withinLocationCountry withinLocationHigherGeography withinLocationType
1 0.12975 -59.65745 Trombetas/Mapuera Brasil Brazil Brasil > Trombetas/Mapuera Território Indígena
2 0.12975 -59.65745 Bioma Amazônia Brasil Brazil Brasil > Bioma Amazônia Ambiental
3 0.12975 -59.65745 Amazonia World Amazonia Ambiental
4 0.12975 -59.65745 Urucará Amazonas Brazil Brasil > Amazonas > Urucará Município
5 1.77469 -59.77757 Jacamim Roraima Brazil Brasil > Roraima > Jacamim Território Indígena
6 1.77469 -59.77757 Bioma Amazônia Brasil Brazil Brasil > Bioma Amazônia Ambiental
withinLocationID withinLocationTypeAdmLevel searchObs
1 6393 98 NA
2 6583 97 NA
3 16597 97 NA
4 1570 8 NA
5 6121 98 NA
6 6583 97 NA
Getting Individual Data
See GET API Individual Endpoint for the full list of search parameters and response fields.
library(opendatabio)
base_url = "https://opendb.inpa.gov.br/api"
token = "YOUR TOKEN HERE"
# Set the connection configuration
cfg = odb_config(base_url = base_url, token = token)
# DIRECT DOWNLOAD – if you want to download a small amount of data
inds = odb_get_individuals(params = list(limit = 100), odb_cfg = cfg)
# PREPARE FILE ON SERVER – if your query will return a large number of records
# Download all records you have access to or public ones
# Save the process, since the result is likely to be large
jobid = odb_get_individuals(params = list(save_job = TRUE), odb_cfg = cfg)
# Check the status of the job
odb_get_jobs(params = list(id = jobid$job_id), odb_cfg = cfg)
# When it finishes, get the data here (or download the file via the web interface)
all_inds = odb_get_jobs(params = list(id = jobid$job_id), odb_cfg = cfg)
# FETCHING SPECIFIC DATA
# All individuals identified as taxon X
params = list(taxon = "Licaria cannela tenuicarpa")
licarias = odb_get_individuals(params = params, odb_cfg = cfg)
# All individuals identified as taxon X or its descendants
params = list(taxon_root = "Licaria")
licarias = odb_get_individuals(params = params, odb_cfg = cfg)
# All individuals from dataset X
params = list(dataset = "MyDataset name or id")
inds = odb_get_individuals(params = params, odb_cfg = cfg)
# Or use save_job above if the dataset is large
# You can view the list of available datasets
datasets = odb_get_datasets(odb_cfg = cfg)
Getting Measurements
See GET API Measurement Endpoint for the complete list of query parameter options and response fields.
Use the odb_get_measurements function.
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token="YOUR TOKEN HERE"
#establishes the connection configuration
cfg = odb_config(base_url=base_url, token = token)
#100 first measurements of the dataset X with id=10
measurements = odb_get_measurements(params=list(dataset=10,limit=100),odb_cfg=cfg)
#100 first measurements of the dataset X with id=10 for the variable whose export_name is treeDbh
measurements = odb_get_measurements(params=list(trait="treeDbh",dataset=10,limit=100),odb_cfg=cfg)
#Measurements of the dataset X with id=10 for the variable whose export_name is treeDbh
#only for Lauraceae
measurements = odb_get_measurements(params=list(trait="treeDbh",dataset=10,taxon_root="Lauraceae"),odb_cfg=cfg)
#linking data of individuals measurements
laurels = odb_get_individuals(params=list(dataset=10,taxon_root="Lauraceae"),odb_cfg=cfg)
filter = grep("Individu",measurements$measured_type) #optional, depends on what is in measurements
g = match(measurements$measured_id[filter],laurels$id)
measurements$location = NA
measurements$location[filter] = laurels$locationName[g]
Getting Media
See GET API Media Endpoint for the complete list of query parameter options and response fields.
Use the odb_get_media function from the R package.
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token="YOUR TOKEN HERE"
#set the connection configuration
cfg = odb_config(base_url=base_url, token = token)
#the first 50 media files of a dataset that has images
imgs = odb_get_media(params=list(dataset=97,limit=50),odb_cfg=cfg)
#see this metadata
head(imgs)
#from this metadata, download the media files
#create a function for this:
getImagesByURL <- function(url,downloadFolder='img') {
dir.create(downloadFolder,showWarnings = F)
fn = strsplit(url,"\\/")[[1]]
fn = fn[length(fn)]
nname = paste(downloadFolder,fn,sep="/")
img = httr::GET(url=url)
writeBin(httr::content(img, "raw"), nname)
}
#use the function to download images to a folder
sapply(imgs$file_url,getImagesByURL,downloadFolder='testeImgsFromOdb')
Getting Voucher Data
See GET API Voucher Endpoint for the full list of search parameter options and response fields.
Follow the example above, but use the odb_get_vouchers function.
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token="YOUR TOKEN HERE"
#establishes the connection configuration
cfg = odb_config(base_url=base_url, token = token)
#first 100 vouchers registered in a biocollection
vouchers = odb_get_vouchers(params=list(biocollection="INPA",limit=100),odb_cfg=cfg)
#vouchers in location x (id, or name, as registered in the database)
vouchers = odb_get_vouchers(params=list(location="Reserva Florestal Adolpho Ducke, Parcela PDBFF-100ha",limit=100),odb_cfg=cfg)
6.2 - Import data with R
The Opendatabio-R package was created to allow users to interact with an OpenDataBio server, to both obtain (GET) data or to import (POST) data into the database. This tutorial is a basic example of how to import data.
Set up the connection
- Set up the connection to the OpenDataBio server using the
odb_config()function. The most important parameters for this function arebase_url, which should point to the API url for your OpenDataBio server, andtoken, which is the access token used to authenticate your user. - The
tokenis mandatory for importing data. - Your token is avaliable in your profile in the web interface
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ"
#create a config object
cfg = odb_config(base_url=base_url, token = token)
#test connection
odb_test(cfg)
Importing data (POST API)
Check the API Quick-Reference for a full list of POST endpoints and link to details.
OpenDataBio-R import functions
All import functions have the same signature: the first argument is a data.frame with data to be imported, and the second parameter is a configuration object generated by odb_config.
When writing an import request, check the POST API docs in order to understand which columns can be declared in the data.frame.
All import functions return a job id, which can be used to check if the job is still running, if it ended with success or if it encountered an error. This job id can be used in the functions odb_get_jobs(), odb_get_affected_ids() and odb_get_log(), to find details about the job, which (if any) were the IDs of the successfully imported objects, and the full log of the job. You may also see the log in your user jobs list in the web interface.
Order is Important - correctly importing data may depend on already registered records. For example, importing an Individual with a Taxon identity requires the taxon name registered in the database, so you would first validate your taxon list using the GET API and then import the records for the Individuals.
You may also Validate geographic coordinates, detecting locations registered in an OpenDataBio database for point geometries.
Working with dates and incomplete dates
For Individuals, Vouchers and Identifications you may use incomplete dates.
The date format used in OpenDataBio is YYY-MM-DD (year - month - day), so a valid entry would be 2018-05-28.
Particularly in historical data, the exact day (or month) may not be known, so you can substitute this fields with NA: ‘1979-05-NA’ means “an unknown day, in May 1979”, and ‘1979-NA-NA’ means “unknown day and month, 1979”. You may not add a date for which you have only the day, but can if you have only the month if is actually meaningful in some way.
6.2.1 - Import Locations
OpenDataBio is distributed with a seed location dataset for Brazil, which includes state, municipality, federal conservation units, indigenous lands and the major biomes.
Working with spatial data is a very delicate area, so we have attempted to make the workflow for inserting locations as easy as possible.
If you want to upload administrative boundaries for a country, you may also just download a geojson file from OSM-Boundaries and upload it through the web interface directly. Or use the GADM repository exemplified below.
Importation is straightforward, but the main issues to keep in mind:
- OpenDataBio stores the geometries of locations using Well-known text (WKT) representation.
- Locations are hierarchical, so a location SHOULD lie completely within its parent location. The importation method will try to detect the parent locations based on its geometry. So you do not need to inform a parent. However, sometimes the parent and child locations share a border or have minor differences that prevent to be detected. Therefore, if this importation fail to place the location where you expected, you may update or import informing the correct parent. When you inform the parent a second check will be performed adding a buffer to the parent location, and should solve the issue.
- Country borders can be imported without parent detection or definition, and marine records may be linked to a parent even if they are not contained by the parent polygon. This requires a specific field specification and should be used only in such cases as it is a possible source of misplacement, but give such flexibility)
- Standardize the geometry to a common projection of use in the system. Strongly recommended to use EPSG:4326 WGS84., for standardization;
- Consider, uploading your political administrative polygons before adding specific POINT, PLOTS or TRANSECTS;
- Conservation Units, Indigenous Territories and Environmental layers may be added as locations and will be treated as special case as some of these locations span different administrative locations. So a POINT, PLOT or TRANSECT location may belong to a UC, a TI and many Environmental layers if these are stored in the database. These related locations, like the political parent, are auto-detected from the location geometry.
Check the POST Locations API docs in order to understand which columns can be declared when importing locations.
Adm_level defines the location type
The administrative level (adm_level) of a location is a number:
- 2 for country; 3 to 10 as other as ‘administrative areas’, following OpenStreeMap convention to facilitate external data importation and local translations (TO BE IMPLEMENTED). So, for Brazil, codes are (States=4, Municipalities=8);
- 999 for ‘POINT’ locations like GPS waypoints;
- 101 for transects
- 100 is the code for plots and subplots;
- 99 is the code for Conservation Units
- 98 for Indigenous Territories
- 97 for Environmental polygons (e.g. Floresta Ombrofila Densa, or Bioma Amazônia)
Importing spatial polygons
GADM Administrative boundaries
Administrative boundaries may also be imported without leaving R, getting data from GDAM and using the odb_import* functions
library(raster)
library(opendatabio)
#download GADM administrative areas for a country
#get country codes
crtcodes = getData('ISO3')
bra = crtcodes[crtcodes$NAME%in%"Brazil",]
#define a path where to save the downloaded spatial data
path = "GADMS"
dir.create(path,showWarnings = F)
#the number of admin_levels in each country varies
#get all levels that exists into your computer
runit =T
level = 0
while(runit) {
ocrt <- try(getData('GADM', country=bra, level=level,path=path),silent=T)
if (class(ocrt)=="try-error") {
runit = FALSE
}
level = level+1
}
#read downloaded data and format to odb
files = list.files(path, full.name=T)
locations.to.odb = NULL
for(f in 1:length(files)) {
ocrt <- readRDS(files[f])
#class(ocrt)
#convert the SpatialPolygonsDataFrame to OpenDataBio format
ocrt.odb = opendatabio:::sp_to_df(ocrt) #only for GADM data
locations.to.odb = rbind(locations.to.odb,ocrt.odb)
}
#see without geometry
head(locations.to.odb[,-ncol(locations.to.odb)])
#you may add a note to location
locations.to.odb$notes = paste("Source gdam.org via raster::get_data()",Sys.Date())
#adjust the adm_level to fit the OpenStreeMap categories
ff = as.factor(locations.to.odb$adm_level)
(lv = levels(ff))
levels(ff) = c(2,4,8,9)
locations.to.odb$adm_level = as.vector(ff)
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ"
cfg = odb_config(base_url=base_url, token = token)
odb_import_locations(data=locations.to.odb,odb_cfg=cfg)
#ATTENTION: you may want to check for uniqueness of name+parent rather than just name, as name+parent are unique for locations. You may not save two locations with the same name within the same parent.
A ShapeFile example
library(rgdal)
#read your shape file
path = 'mymaps'
file = 'myshapefile.shp'
layer = gsub(".shp","",file,ignore.case=TRUE)
data = readOGR(dsn=path, layer= layer)
#you may reproject the geometry to standard of your system if needed
data = spTransform(data,CRS=CRS("+proj=longlat +datum=WGS84"))
#convert polygons to WKT geometry representation
library(rgeos)
geom = rgeos::writeWKT(data,byid=TRUE)
#prep import
names = data@data$name #or the column name of the data
shape.to.odb = data.frame(name=names,geom=geom,stringsAsFactors = F)
#need to add the admin_level of these locations
shape.to.odb$admin_level = 2
#and may add parent and note if your want
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ"
cfg = odb_config(base_url=base_url, token = token)
odb_import_locations(data=shape.to.odb,odb_cfg=cfg)
Converting data from KML
#read file as SpatialPolygonDataFrame
file = "myfile.kml"
file.exists(file)
mykml = readOGR(file)
geom = rgeos::writeWKT(mykml,byid=TRUE)
#prep import
names = mykml@data$name #or the column name of the data
to.odb = data.frame(name=names,geom=geom,stringsAsFactors = F)
#need to add the admin_level of these locations
to.odb$admin_level = 2
#and may add parent or any other valid field
#import
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ"
cfg = odb_config(base_url=base_url, token = token)
odb_import_locations(data=to.odb,odb_cfg=cfg)
Import Plots and Subplots
Plots and Transects are special cases within OpenDataBio:
- They may be defined with a Polygon or LineString geometry, respectively;
- Or they may be registered only as POINT locations. In this case OpenDataBio will create the polygon or linestring geometry for you;
- Dimensions (x and y) are stored in meters
- SubPlots are plot locations having a plot location as a parent, and must also have cartesian positions (startX, startY) within the parent location in addition to dimensions. Cartesian position refer to X and Y positions within parent plot and hence MUST be smaller than parent X and Y. And the same is true for Individuals within plots or subplots when they have their own X and Y cartesian coordinates.
- SubPlot is the only location that may be registered without a geographical coordinate or geometry, which will be calculated from the parent plot geometry using the startx and starty values.
Plot and subplot example 01
You need at least a single point geographical coordinate for a location of type PLOT. Geometry (or lat and long) cannot be empty.
#geometry of a plot in Manaus
southWestCorner = c(-59.987747, -3.095764)
northWestCorner = c(-59.987747, -3.094822)
northEastCorner = c(-59.986835,-3.094822)
southEastCorner = c(-59.986835,-3.095764)
geom = rbind(southWestCorner,northWestCorner,northEastCorner,southEastCorner)
library(sp)
geom = Polygon(geom)
geom = Polygons(list(geom), ID = 1)
geom = SpatialPolygons(list(geom))
library(rgeos)
geom = writeWKT(geom)
to.odb = data.frame(name='A 1ha example plot',x=100,y=100,notes='a fake plot',geom=geom, adm_level = 100,stringsAsFactors=F)
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ"
cfg = odb_config(base_url=base_url, token = token)
odb_import_locations(data=to.odb,odb_cfg=cfg)
Wait a few seconds, and then import subplots to this plot.
#import 20x20m subplots to the plot above without indicating a geometry.
#SubPlot is the only location type that does not require the specification of a geometry or coordinates,
#but it requires specification of startx and starty relative position coordinates within parent plot
#OpenDataBio will use subplot position values to calculate its geographical coordinates based on parent geometry
(parent = odb_get_locations(params = list(name='A 1ha example plot',fields='id,name',adm_level=100),odb_cfg = cfg))
sub1 = data.frame(name='sub plot 40x40',parent=parent$id,x=20,y=20,adm_level=100,startx=40,starty=40,stringsAsFactors=F)
sub2 = data.frame(name='sub plot 0x0',parent=parent$id,x=20,y=20,adm_level=100,startx=0,starty=0,stringsAsFactors=F)
sub3 = data.frame(name='sub plot 80x80',parent=parent$id,x=20,y=20,adm_level=100,startx=80,starty=80,stringsAsFactors=F)
dt = rbind(sub1,sub2,sub3)
#import
odb_import_locations(data=dt,odb_cfg=cfg)
Screen captures of imported plots
Below screen captures for the locations imported with the code above
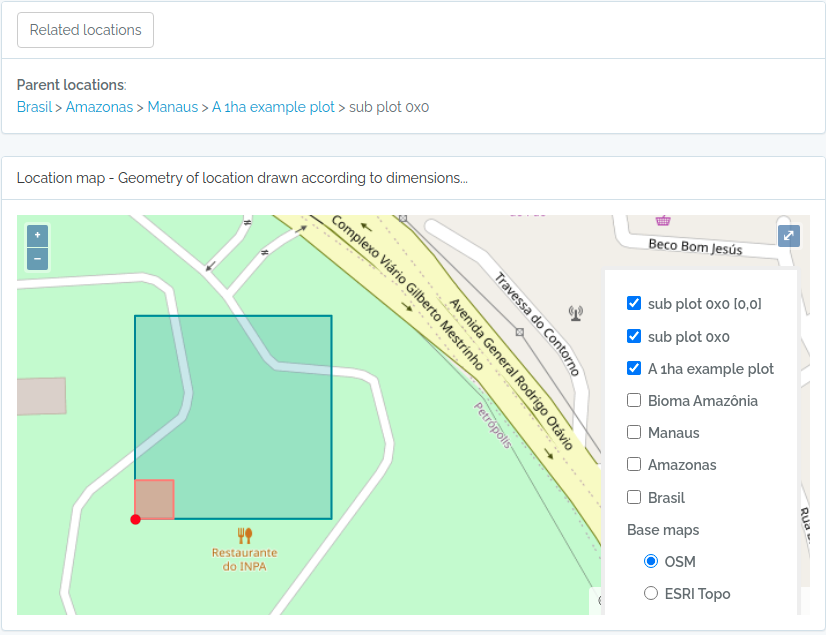
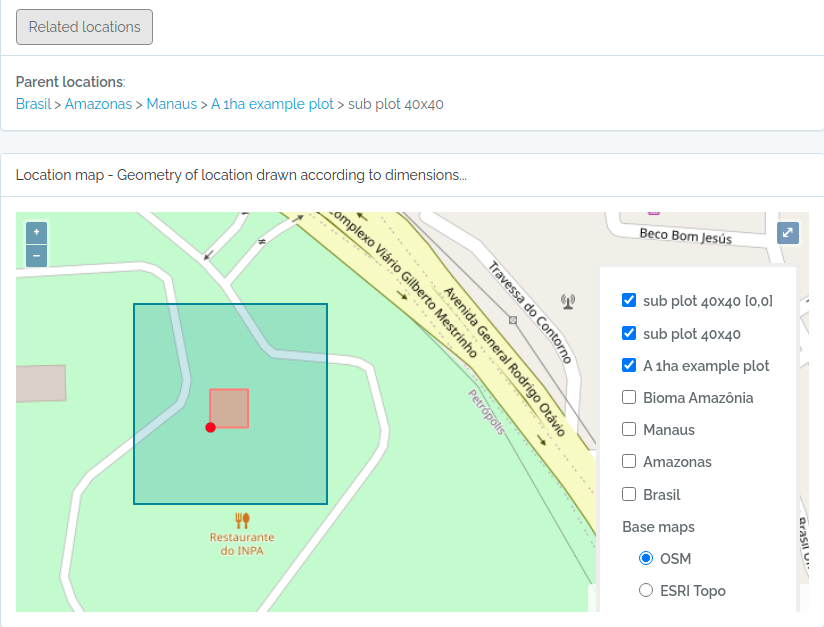
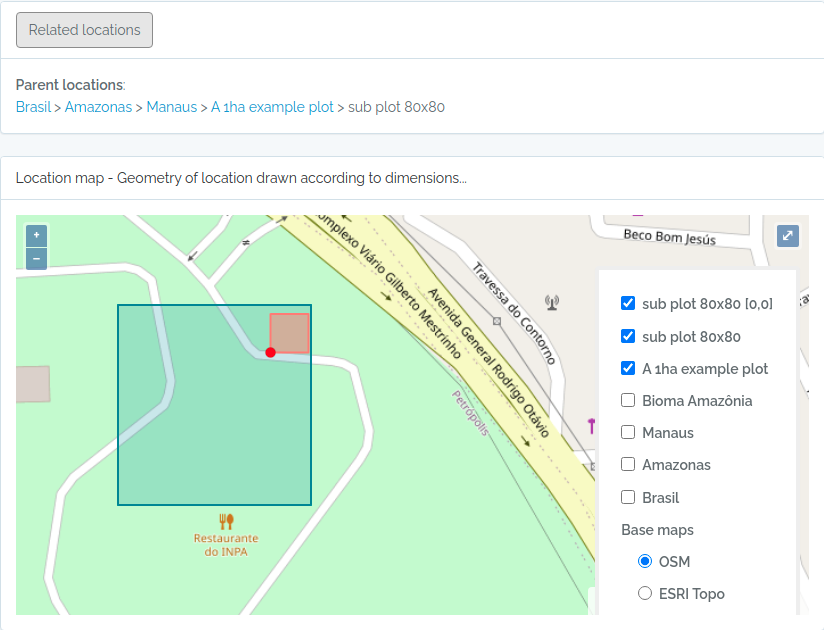
Plot and subplot example 02
Import a plot and subplots having only:
- a single point coordinate
- an azimuth or angle of the plot direction
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ"
cfg = odb_config(base_url=base_url, token = token)
#the plot
geom = "POINT(-59.973841 -2.929822)"
to.odb = data.frame(name='Example Point PLOT',x=100, y=100, azimuth=45,notes='OpenDataBio point plot example',geom=geom, adm_level = 100,stringsAsFactors=F)
odb_import_locations(data=to.odb,odb_cfg=cfg)
#define 20x20 subplots cartesian coordinates
x = seq(0,80,by=20)
xx = rep(x,length(x))
yy = rep(x,each=length(x))
names = paste(xx,yy,sep="x")
#import these subplots without having a geometry, but specifying the parent plot location
parent = odb_get_locations(params = list(name='Example Point PLOT',adm_level=100),odb_cfg = cfg)
to.odb = data.frame(name=names,startx=xx,starty=yy,x=20,y=20,notes="OpenDataBio 20x20 subplots example",adm_level=100,parent=parent$id)
odb_import_locations(data=to.odb,odb_cfg=cfg)
#get the imported plot locations and plot them using the root parameter
locais = odb_get_locations(params=list(root=parent$id),odb_cfg = cfg)
locais[,c('id','locationName','parentName')]
colnames(locais)
for(i in 1:nrow(locais)) {
geom = readWKT(locais$footprintWKT[i])
if (i==1) {
plot(geom,main=locais$locationName[i],cex.main=0.8,col='yellow')
axis(side=1,cex.axis=0.7)
axis(side=2,cex.axis=0.7,las=2)
} else {
plot(geom,add=T,border='red')
}
}
The figure generated above:
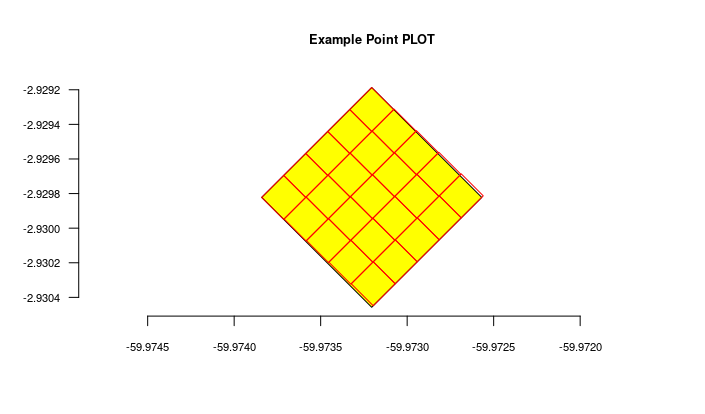
Import transects
This code will import two transects, when defined by a LINESTRING geometry, the other only by a point geometry. See figures below for the imported result
#geometry of transect in Manaus
#read trail from a kml file
#library(rgdal)
#file = "acariquara.kml"
#file.exists(file)
#mykml = readOGR(file)
#library(rgeos)
#geom = rgeos::writeWKT(mykml,byid=TRUE)
#above will output:
geom = "LINESTRING (-59.9616459699999993 -3.0803612500000002, -59.9617394400000023 -3.0805952900000002, -59.9618530300000003 -3.0807376099999999, -59.9621049400000032 -3.0808563200000001, -59.9621949100000009 -3.0809758500000002, -59.9621587999999974 -3.0812666800000001, -59.9621092399999966 -3.0815010400000000, -59.9620656999999966 -3.0816403499999998, -59.9620170600000009 -3.0818584699999998, -59.9620740699999999 -3.0819864099999998)";
#prep data frame
#the y value refer to a buffer in meters applied to the trail
#y is used to validate the insertion of related individuals
to.odb = data.frame(name='A trail-transect example',y=20, notes='OpenDataBio transect example',geom=geom, adm_level = 101,stringsAsFactors=F)
#import
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ"
cfg = odb_config(base_url=base_url, token = token)
odb_import_locations(data=to.odb,odb_cfg=cfg)
#NOW IMPORT A SECOND TRANSECT WITHOUT POINT GEOMETRY
#then you need to inform the x value, which is the transect length
#ODB will map this transect oriented by the azimuth paramater (south in the example below)
#point geometry = start point
geom = "POINT(-59.973841 -2.929822)"
to.odb = data.frame(name='A transect point geometry',x=300, y=20, azimuth=180,notes='OpenDataBio point transect example',geom=geom, adm_level = 101,stringsAsFactors=F)
odb_import_locations(data=to.odb,odb_cfg=cfg)
locais = odb_get_locations(params=list(adm_level=101),odb_cfg = cfg)
locais[,c('id','locationName','parentName','levelName')]
The code above will result in the following two locations:
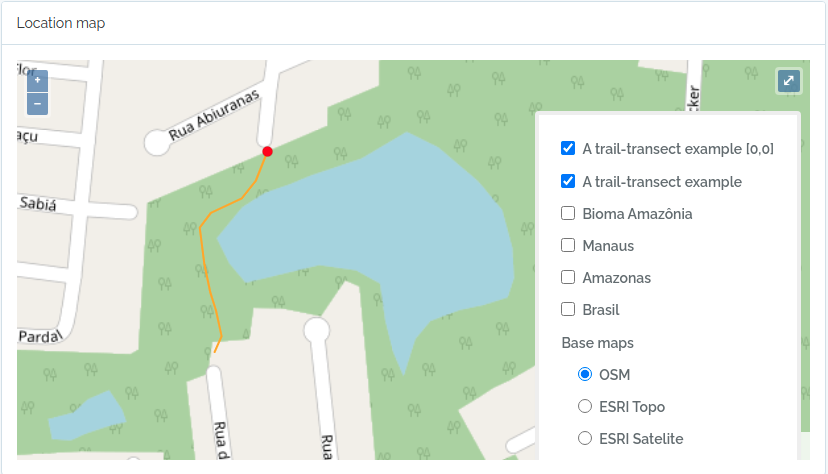
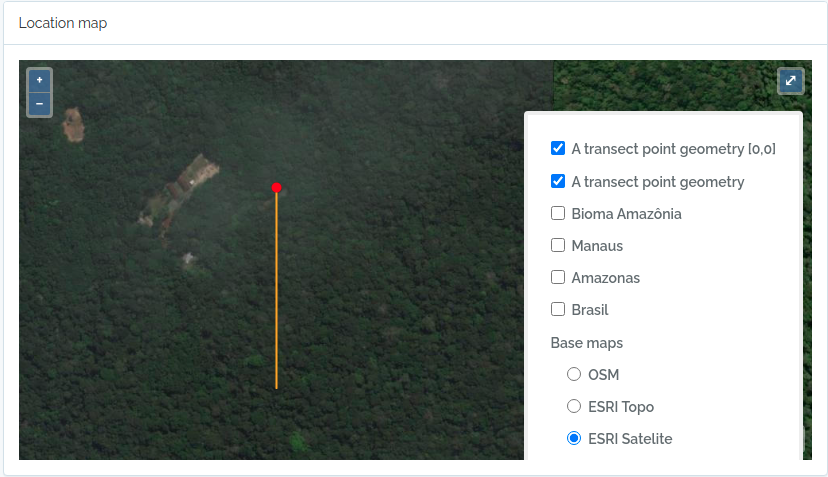
6.2.2 - Import BibReferences
#your connection
library(opendatabio)
base_url="http://localhost/opendatabio/api"
token = “YOUR TOKEN HERE”
cfg = odb_config(base_url=base_url, token = token)
odb_test(cfg)
#read the bibliographic references in R
library(rbibutils)
bibs = readBib(file="yourFileWithReferences.bib")
formatbib <- function(x) {
con <- textConnection("bibref", "w")
writeBib(x,con=con)
bibref = paste(bibref,collapse = " ")
close(con)
return(bibref)
}
#prepare to import to odb
bibtexts = sapply(bibs,formatbib)
data = data.frame(bibtex=bibtexts,standardize=1,stringsAsFactors = F)
#importa
jobid = odb_import_bibreferences(data,odb_cfg = cfg)
#waiting for completion
odb_get_jobs(params=list(id=jobid$id),odb_cfg = cfg)
#get the import log
dt = odb_get_affected_ids(job_id=jobid$id,odb_cfg = cfg)
6.2.3 - Import Vernacular
library(opendatabio)
base_url="http://localhost/opendatabio/api"
token="YOUR TOKEN HERE"
cfg = odb_config(base_url=base_url, token = token)
odb_test(cfg)
#generate fake data for testing
name = c('pau rosa',"casca preciosa")
taxons = c("Aniba roseaodora,Aniba panurensis,Aniba parvifolia","Aniba canelilla")
#get the id of some individuals
inds = odb_get_individuals(params=list(taxon="Aniba roseaodora,Aniba panurensis,Aniba parvifolia",fields='id,scientificName',limit=10),odb_cfg = cfg)
individuals=c(paste(inds$id,collapse = ","),NA)
#idiomas = odb_get_languages(odb_cfg = cfg)
language= c('pt-br','en')
#create a data.frame with this information
verna = data.frame(name,taxons,language,taxons,individuals)
#citations (just generate a data.frame for each vernacular)
umaCitatcao = data.frame(
citation='This would be the cited text',
bibreference="Riberiroetal1999FloraDucke", #bibkey or the id
type='generic', #can be: generic, use, etymology
notes='my observations about this citation')
#add a list-type column
verna$citations = list(oneCitation,NA)
#import to opendatabio
odb_import_vernaculars(verna,odb_cfg = cfg)
6.2.4 - Import Media
- You basically need the media files in a single folder and a table containing the attributes for each media file;
- See the POST Media API docs to understand which columns can be declared when importing Media. Only 3 columns are mandatory in the attribute table:
filename,object_type, andobject_id, but it is important to include additional information about the file, such as date, authors, title, usage license, dataset, and also Tags. - Consider adding at least one keyword (Tag) for each media file.
Importing Media via API
The method below can be done either through the API using the R package or through the web interface. Test and use whichever is faster to upload the image files.
library(opendatabio)
base_url = "https://opendb.inpa.gov.br/api"
token = "GZ1iXcmRvIFQ"
cfg = odb_config(base_url = base_url)
# path to the folder where the images are stored
folder = 'imagesParaOdb'
# list the file names
filenames = dir(folder, full.names = FALSE)
# read the attribute table
attributes = read.table('arquivoAtributos.csv', sep = ',', header = TRUE, as.is = TRUE, na.strings = c("", "NA", "-"))
# are all files listed in the attribute table?
print(paste(sum(filenames %in% attributes$filename), "of", length(filenames), "files are listed in the attribute table"))
# import to ODB
odb_upload_media_zip(folder = folder, attribute_table = attributes, odb_cfg = cfg)
6.2.5 - Import Taxons
- For a successful importation of published names, you just need to provide a
nameattribute, being the fullname for species and infraspecies (e.g. Licaria canella tenuicarpa, Ocotea delicata), same form asGBIF's canonicalName, i.e. do not include infraspecific ranks; - No need to import parent taxons for published names, OpenDataBio does that for you. The import process will use the
nameyou informed to retrieve taxon info from a nomenclature repository: currently Tropicos, GBIF BackBone Taxonomy and ZOOBANK. A fuzzy search will be performed on the GBIF service if needed. If the taxon name is found, it will automatically get the needed information and also all parent taxa (or accepted names if its tagged as a synonym) as needed and will store in the database for you. If your name is invalid, i.e. it is a synonym of a another name, it will be imported but tagged as invalid. - If you inform
parentand it is different from the one detected by the API the informed parent has preference. - For unpublished species
level,parent, and apersonorauthor_idmust also be provided; - The external API will use a fuzzy query while searching GBIF if it does not find an exact match, allowing to detect even when misspelled. However, spelling errors in names may prevent a taxon from being stored.
- Taxonomy is hierarchical and the structure within the OpenDataBio may be treelike. So, you may add, if you want, any node of the tree of life into the Taxon table. For these you need to use the
Cladetaxon level. But note, this may not be relevant if you are not going link data to such clade names. - Because the nomenclature check is time consuming, be patient.
OpenDataBio is distributed with a Taxon seed table, that include the root of all kingdoms and some high-level nodes for Plants, i.e. the APWeb Order level tree for Angiosperms.
A simple published name example
The scripts below were tested on top of the OpenDataBio Seed Taxon table, which contains only down to the order level for Angiosperms.
In the taxons table, families Moraceae, Lauraceae and Solanaceae were not yet registered:
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
cfg = odb_config(base_url=base_url)
exists = odb_get_taxons(params=list(root="Moraceae,Lauraceae,Solanaceae"),odb_cfg=cfg)
Returned:
data frame with 0 columns and 0 rows
Now import some species and one infraspecies for the families above, specifying their fullname (canonicalName):
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ"
cfg = odb_config(base_url=base_url, token = token)
spp = c("Ficus schultesii", "Ocotea guianensis","Duckeodendron cestroides","Licaria canella tenuicarpa")
splist = data.frame(name=spp)
odb_import_taxons(splist, odb_cfg=cfg)
Now check with the same code for taxons under those children:
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
cfg = odb_config(base_url=base_url)
exists = odb_get_taxons(params=list(root="Moraceae,Lauraceae,Chrysobalanaceae"),odb_cfg=cfg)
head(exists[,c('id','scientificName', 'taxonRank','taxonomicStatus','parentNameUsage')])
Which will return:
id scientificName taxonRank taxonomicStatus parentName
1 252 Moraceae Family accepted Rosales
2 253 Ficus Genus accepted Moraceae
3 254 Ficus schultesii Species accepted Ficus
4 258 Solanaceae Family accepted Solanales
5 259 Duckeodendron Genus accepted Solanaceae
6 260 Duckeodendron cestroides Species accepted Duckeodendron
7 255 Lauraceae Family accepted Laurales
8 256 Ocotea Genus accepted Lauraceae
9 257 Ocotea guianensis Species accepted Ocotea
10 261 Licaria Genus accepted Lauraceae
11 262 Licaria canella Species accepted Licaria
12 263 Licaria canella subsp. tenuicarpa Subspecies accepted Licaria canella
Note that although we specified only the species and infraspecies names, the API imported also all the needed parent hierarchy up to family, because orders where already registered.
An invalid published name example
The name Licania octandra pallida (Chrysobalanaceae) has been recently turned into a synonym of Leptobalanus octandrus pallidus.
The script below exemplify what happens in such cases.
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ"
cfg = odb_config(base_url=base_url, token = token)
#lets check
exists = odb_get_taxons(params=list(root="Chrysobalanaceae"),odb_cfg=cfg)
exists
#in this test returns an empty data frame
#data frame with 0 columns and 0 rows
#now import
spp = c("Licania octandra pallida")
splist = data.frame(name=spp)
odb_import_taxons(splist, odb_cfg=cfg)
#see the results
exists = odb_get_taxons(params=list(root="Chrysobalanaceae"),odb_cfg=cfg)
exists[,c('id','scientificName', 'taxonRank','taxonomicStatus','parentName')]
Which will return:
id scientificName taxonRank taxonomicStatus parentName
1 264 Chrysobalanaceae Family accepted Malpighiales
2 265 Leptobalanus Genus accepted Chrysobalanaceae
3 267 Leptobalanus octandrus Species accepted Leptobalanus
4 269 Leptobalanus octandrus subsp. pallidus Subspecies accepted Leptobalanus octandrus
5 266 Licania Genus accepted Chrysobalanaceae
6 268 Licania octandra Species invalid Licania
7 270 Licania octandra subsp. pallida Subspecies invalid Licania octandra
Note that although we specified only one infraspecies name, the API imported also all the needed parent hierarchy up to family, and because the name is invalid it also imported the acceptedName for this infraspecies and its parents.
An unpublished species or morphotype
It is common to have unpublished local species names (morphotypes) for plants in plots, or yet to be published taxonomic work. Unpublished designation are project specific and therefore MUST also provide an author as different projects may use the same ‘sp.1’ or ‘sp.A’ code for their unpublished taxons.
You may link an unpublished name as any taxon level and do not need to use genus+species logic to assign a morphotype for which the genus or upper level taxonomy is undefined. For example, you may store a ‘species’ level with name ‘Indet sp.1’ and parent_name ‘Laurales’, if the lowest level formal determination you have is the order level. In this example, there is no need to store a Indet genus and Indet family taxons just to account for this unidentified morphotype.
##assign an unpublished name for which you only know belongs to the Angiosperms and you have this node in the Taxon table already
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
cfg = odb_config(base_url=base_url)
#check that angiosperms exist
odb_get_taxons(params=list(name='Angiosperms'),odb_cfg = cfg)
#if it is there, start creating a data.frame to import
to.odb = data.frame(name='Morphotype sp.1', parent='Angiosperms', stringsAsFactors=F)
#get species level numeric code
to.odb$level=odb_taxonLevelCodes('species')
#you must provide an author that is a Person in the Person table. Get from server
odb.persons = odb_get_persons(params=list(search='João Batista da Silva'),odb_cfg=cfg)
#found
head(odb.persons)
#add the author_id to the data.frame
#NOTE it is not author, but author_id or person)
#this makes odb understand it is an unpublished name
to.odb$author_id = odb.persons$id
#import
token ="GZ1iXcmRvIFQ" #this must be your token not this value
cfg = odb_config(base_url=base_url, token = token)
odb_import_taxons(to.odb,odb_cfg = cfg)
Check the imported record:
exists = odb_get_taxons(params=list(name='Morphotype sp.1'),odb_cfg = cfg)
exists[,c('id','scientificName', 'taxonRank','taxonomicStatus','parentName','scientificNameAuthorship')]
Some columns for the imported record:
id scientificName taxonRank taxonomicStatus parentName scientificNameAuthorship
1 276 Morphotype sp.1 Species unpublished Angiosperms João Batista da Silva - Silva, J.B.D.
Import a published clade
You may add a clade Taxon and may reference its publication using the bibkey entry. So, it is possible to actually store all relevant nodes of any phylogeny in the Taxon hierarchy.
#parent must be stored already
odb_get_taxons(params=list(name='Pagamea'),odb_cfg = cfg)
#define clade Taxon
to.odb = data.frame(name='Guianensis core', parent_name='Pagamea', stringsAsFactors=F)
to.odb$level = odb_taxonLevelCodes('clade')
#add a reference to the publication where it is published
#import bib reference to database beforehand
odb_get_bibreferences(params(bibkey='prataetal2018'),odb_cfg=cfg)
to.odb$bibkey = 'prataetal2018'
#then add valid species names as children of this clade instead of the genus level
children = data.frame(name = c('Pagamea guianensis','Pagamea angustifolia','Pagamea puberula'),stringsAsFactors=F)
children$parent_name = 'Guianensis core'
children$level = odb_taxonLevelCodes('species')
children$bibkey = NA
#merge
to.odb = rbind(to.odb,children)
#import
odb_import_taxons(to.odb,odb_cfg = cfg)
6.2.6 - Import Persons
Check the POST Persons API docs in order to understand which columns can be declared when importing Persons.
It is recommended you use the web interface, as it will warn you in case the person you want to register has a similar, likely the same, person already registered. The API will only check for identical Abbreviations, which is the single restriction of the Person class. Abbreviations are unique and duplications are not allowed. This does not prevent data downloaded from repositories to have different abbreviations or full name for the same person. So, you should standardize secondary data before importing into the server to minimize such common errors.
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ"
cfg = odb_config(base_url=base_url, token = token)
one = data.frame(full_name='Adolpho Ducke',abbreviation='DUCKE, A.',notes='Grande botânico da Amazônia',stringsAsFactors = F)
two = data.frame(full_name='Michael John Gilbert Hopkins',abbreviation='HOPKINKS, M.J.G.',notes='Curador herbário INPA',stringsAsFactors = F)
to.odb= rbind(one,two)
odb_import_persons(to.odb,odb_cfg=cfg)
#may also add an email entry if you have one
Get the data
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
cfg = odb_config(base_url=base_url)
persons = odb_get_persons(odb_cfg=cfg)
persons = persons[order(persons$id,decreasing = T),]
head(persons,2)
Will output:
id full_name abbreviation email institution notes
613 1582 Michael John Gilbert Hopkins HOPKINKS, M.J.G. <NA> NA Curador herbário INPA
373 1581 Adolpho Ducke DUCKE, A. <NA> NA Grande botânico da Amazônia
6.2.7 - Import Traits
- Strongly recommended you use the web interface to enter traits one by one. Use batch imports only if you have too many.
- Before importing any trait, make sure the variable is not already registered in the system with a different name. Preventing duplication is important as the Trait class is shared among all users of an OpenDataBio installation. This also means that once a trait is used for Measurements, you may only alter the trait’s definition, for example, a category name, if you were the only user entering measurements for the trait. Otherwise, somebody else used that definition and you will not be able to alter it.
- Export_name must be unique in a single ODB installation. This name is used when exporting data and on forms, and you should consider making it as short and informative (of definition) as possible. Also, API will validate the export name - must be camelCase, PascalCase or snake_case, and you cannot add spaces or any special character (accents). Ex. ‘dbh’,‘dbhPom’, ’leafLength’, ‘LeafLength’ or ’leaf_length’, respectively.
Traits can be imported using odb_import_traits().
Read carefully the Traits POST API.
Traits types
See odb_traitTypeCodes() for possible trait types.
Trait name and categories User translations
Fields name and description could be one of following:
- using the Language code as keys:
list("en" = "Diameter at Breast Height","pt-br" ="Diâmetro a Altura do Peito") - or using the Language names as keys:
list("English" ="Diameter at Breast Height","Portuguese" ="Diâmetro a Altura do Peito").
Field categories must include for each category+rank+lang the following fields: lang=mixed - required, the id, code or name of the language of the translationname=string - required, the translated category name required (name+rank+lang must be unique)rank=number - required, rank is important to indicate the same category across languages, and defines ordinal traits;description=string - optional for categories, a definition of the category.
This may be formatted as a data.frame and placed in the categories column of another data.frame:
data.frame(
rbind(
c("lang"="en","rank"=1,"name"="small","description"="smaller than 1 cm"),
c("lang"="pt-br","rank"=1,"name"="pequeno","description"="menor que 1 cm"),
c("lang"="en","rank"=2,"name"="big","description"="bigger than 10 cm"),
c("lang"="pt-br","rank"=2,"name"="grande","description"="maior que 10 cm")
),
stringsAsFactors=FALSE
)
Quantitative trait example
For quantitative traits for either integers or real values (types 0 or 1).
odb_traitTypeCodes()
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ" #this must be your token not this value
cfg = odb_config(base_url=base_url, token = token)
#do this first to build a correct data.frame as it will include translations list
to.odb = data.frame(type=1,export_name = "dbh", unit='centimeters',stringsAsFactors = F)
#add translations (note double list)
#format is language_id = translation (and the column be a list with the translation lists)
to.odb$name[[1]]= list('1' = 'Diameter at breast height', '2' = 'Diâmetro à altura do peito')
to.odb$description[[1]]= list('1' = 'Stem diameter measured at 1.3m height','2' = 'Diâmetro do tronco medido à 1.3m de altura')
#measurement validations
to.odb$range_min = 10 #this will restrict the minimum measurement value allowed in the trait
to.odb$range_max = 400 #this will restrict the maximum value
#measurements can be linked to (classes concatenated by , or a list)
to.odb$objects = "Individual,Voucher,Taxon" #makes no sense link such measurements to Locations
to.odb$notes = 'this is quantitative trait example'
#import
odb_import_traits(to.odb,odb_cfg=cfg)
Categorical trait example
- Must include categories. The only difference between ordinal and categorical traits is that ordinal categories will have a rank and the rank will be inferred from the order the categories are informed during importation. Note that ordinal traits are semi-quantitative and so, if you have categories ask yourself whether they are not really ordinal and register accordingly.
- Like the Trait name and description, categories may also have different language translations, and you SHOULD enter the translations for the languages available (
odb_get_languages()) in the server, so the Trait will be accessible in all languages. English is mandatory, so at least the English name must be informed. Categories may have a description associated, but sometimes the category name is self explanatory, so descriptions of categories are not mandatory.
odb_traitTypeCodes()
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ" #this must be your token not this value
cfg = odb_config(base_url=base_url, token = token)
#do this first to build a correct data.frame as it will include translations list
#do this first to build a correct data.frame as it will include translations list
to.odb = data.frame(type=3,export_name = "specimenFertility", stringsAsFactors = F)
#trait name and description
to.odb$name = data.frame("en"="Specimen Fertility","pt-br"="Fertilidade do especímene",stringsAsFactors=F)
to.odb$description = data.frame("en"="Kind of reproductive stage of a collected plant","pt-br"="Estágio reprodutivo de uma amostra de planta coletada",stringsAsFactors=F)
#categories (if your trait is ORDINAL, the add categories in the wanted order here)
categories = data.frame(
rbind(
c('en',1,"Sterile"),
c('pt-br',1,"Estéril"),
c('en',2,"Flowers"),
c('pt-br',2,"Flores"),
c('en',3,"Fruits"),
c('pt-br',3,"Frutos"),
c('en',4,"Flower buds"),
c('pt-br',4,"Botões florais")
),
stringsAsFactors =FALSE
)
colnames(categories) = c("lang","rank","name")
#descriptions not included for categories as they are obvious,
# but you may add a 'description' column to the categories data.frame
#objects for which the trait may be used for
to.odb$objects = "Individual,Voucher"
to.odb$notes = 'a fake note for a multiselection categorical trait'
to.odb$categories = list(categories)
#import
odb_import_traits(to.odb,odb_cfg=cfg)
Link type trait example
Link types are traits that allow you link a Taxon or Voucher as a value measurement to another object. For example, you may conduct a plant inventory for which you have only counts for Taxon associated to a locality. Therefore, you may create a LINK trait, which will allow you to store the count values for any Taxon as measurements for a particular location (POINT, POLYGON). Or you may link such values to Vouchers instead of Taxons if you have a representative specimen for the taxons.
Use the WebInterface.
Text and color traits
Text and color traits require the minimum fields only for trait registration. Text traits allow the storage of textual observations. Color will only allow color codes (see example in Importing Measurements)
odb_traitTypeCodes()
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ" #this must be your token not this value
cfg = odb_config(base_url=base_url, token = token)
to.odb = data.frame(type=5,export_name = "taxonDescription", stringsAsFactors = F)
#trait name and description
to.odb$name = data.frame("en"="Taxonomic descriptions","pt-br"="Descrições taxonômicas",stringsAsFactors=F)
to.odb$description = data.frame("en"="Taxonomic descriptions from the literature","pt-br"="Descrições taxonômicas da literatura",stringsAsFactors=F)
#will only be able to use this trait for a measurment associated with a Taxon
to.odb$objects = "Taxon"
#import
odb_import_traits(to.odb,odb_cfg=cfg)
Spectral traits
Spectral traits are specific to spectral data. You must specify the range of wavenumber values for which you may have absorbance or reflectance data, and the length of the spectra to be stored as measurements to allow validation during input. So, for each range and spacement of the spectral values you have, a different SPECTRAL trait must be created.
Use the WebInterface.
6.2.8 - Import Individuals & Vouchers
- You may import vouchers with the Individual POST endpoint. Use the voucher endpoint only for already registered individuals, else follow below to import individuals and their vouchers at once.
- Individuals may have a self taxonomic identification or a non-self taxonomic identification
- Individuals may have multiple locations, but when registering the individual a single location is needed. You may then import additional locations using the Individual-location POST API.
Individuals can be imported using odb_import_individuals() and vouchers with the odb_import_vouchers().
Read carefully the Individual POST API and the Voucher POST API.
Individual example
Prep data for a single individual basic example, representing a tree in a forest plot location.
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ" #this must be your token not this value
cfg = odb_config(base_url=base_url, token = token)
#the number in the aluminium tag in the forest
to.odb = data.frame(tag='3405.L1', stringsAsFactors=F)
#the collectors (get ids from the server)
(joao = odb_get_persons(params=list(search='joao batista da silva'),odb_cfg=cfg)$id)
(ana = odb_get_persons(params=list(search='ana cristina sega'),odb_cfg=cfg)$id)
#ids concatenated by | pipe
to.odb$collector = paste(joao,ana,sep='|')
#tagged date (lets use an incomplete).
to.odb$date = '2018-07-NA'
#lets place in a Plot location imported with the Location post tutorial
plots = odb_get_locations(params=list(name='A 1ha example plot'),odb_cfg=cfg)
head(plots)
to.odb$location = plots$id
#relative position within parent plot
to.odb$x = 10.4
to.odb$y = 32.5
#or could be
#to.odb$relative_position = paste(x,y,sep=',')
#taxonomic identification
taxon = 'Ocotea guianensis'
#check that exists
(odb_get_taxons(params=list(name='Ocotea guianensis'),odb_cfg=cfg)$id)
#person that identified the individual
to.odb$identifier = odb_get_persons(params=list(search='paulo apostolo'),odb_cfg=cfg)$id
#or you also do to.odb$identifier = "Assunção, P.A.C.L."
#the used form only guarantees the persons is there.
#may add modifers as well [may need to use numeric code instead]
to.odb$modifier = 'cf.'
#or check with to see you spelling is correct
odb_detModifiers()
#and submit the numeric code instaed
to.odb$modifier = 3
#an incomplete identification date
to.odb$identification_date = list(year=2005)
#or to.odb$identification_date = "2005-NA-NA"
Lets import the above record:
odb_import_individuals(to.odb,odb_cfg = cfg)
#lets import this individual
odb_import_individuals(to.odb,odb_cfg = cfg)
#check the job status
odb_get_jobs(params=list(id=130),odb_cfg = cfg)
Ops, I forgot to inform a dataset and my user does not have a default dataset defined.
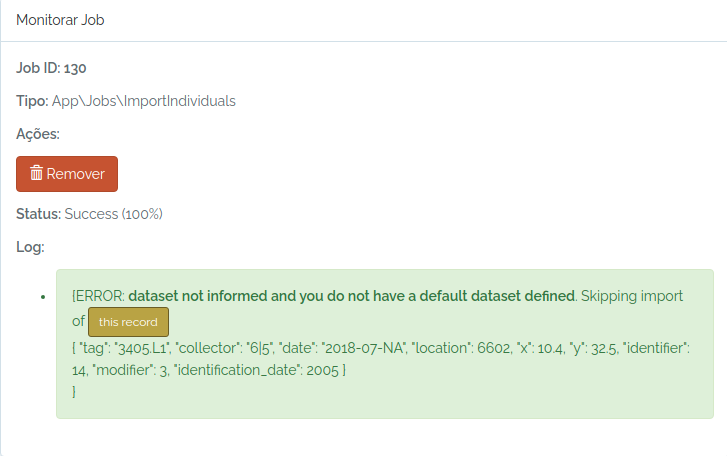
So, I just inform an existing dataset and try again:
dataset = odb_get_datasets(params=list(name="Dataset test"),odb_cfg=cfg)
dataset
to.odb$dataset = dataset$id
odb_import_individuals(to.odb,odb_cfg = cfg)
The individual was imported. The image below shows the individual (yellow dot) mapped in the plot:
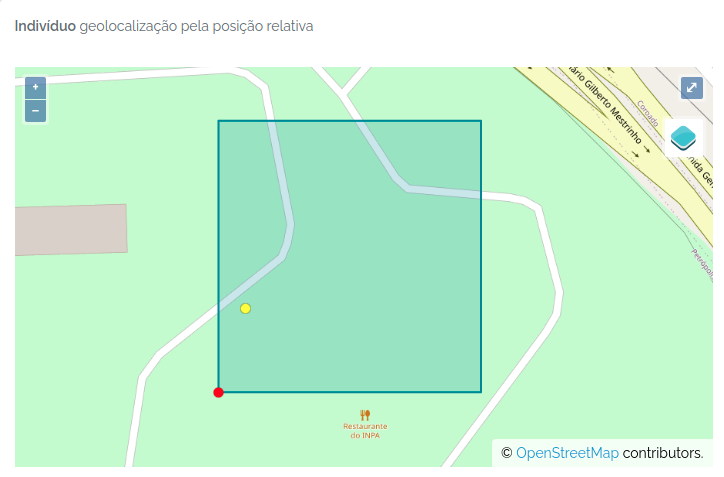
Importing Individuals and Vouchers at once
Individuals are the actual object that has most of the information related to Vouchers, which are samples in a Biocollection. Therefore, you may import an individual record with the specification of one or more vouchers.
#a fake plant record somewhere in the Amazon
aplant = data.frame(taxon="Duckeodendron cestroides", date="2021-09-09", latitude=-2.34, longitude=-59.845,angle=NA,distance=NA, collector="Oliveira, A.A. de|João Batista da Silva", tag="3456-A",dataset=1)
#a fake set of vouchers for this individual
herb = data.frame(biocollection=c("INPA","NY","MO"),biocollection_number=c("12345A","574635","ANOTHER FAKE CODE"),biocollection_type=c(2,3,3))
#add this dataframe to the object
aplant$biocollection = NA
aplant$biocollection = list(herb)
#another fake plant
asecondplant = data.frame(taxon="Ocotea guianensis", date="2021-09-09", latitude=-2.34, longitude=-59.89,angle=240,distance=50, collector="Oliveira, A.A. de|João Batista da Silva", tag="3456",dataset=1)
asecondplant$biocollection = NA
#merge the fake data
to.odb = rbind(aplant,asecondplant)
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ" #this must be your token not this value
cfg = odb_config(base_url=base_url, token = token)
odb_import_individuals(to.odb, odb_cfg=cfg)
Check the imported data
The script above has created records for both the Individual and Voucher model:
#get the imported individuals using a wildcard
inds = odb_get_individuals(params = list(tag='3456*'),odb_cfg = cfg)
inds[,c("basisOfRecord","scientificName","organismID","decimalLatitude","decimalLongitude","higherGeography") ]
Will return:
basisOfRecord scientificName organismID decimalLatitude decimalLongitude higherGeography
1 Organism Ocotea guianensis 3456 - Oliveira - UnnamedPoint_5989234 -2.3402 -59.8904 Brasil | Amazonas | Rio Preto da Eva
2 Organism Duckeodendron cestroides 3456-A - Oliveira - UnnamedPoint_5989234 -2.3400 -59.8900 Brasil | Amazonas | Rio Preto da Eva
And the vouchers:
#get the vouchers imported with the first plant data
vouchers = odb_get_vouchers(params = list(individual=inds$id),odb_cfg = cfg)
vouchers[,c("basisOfRecord","scientificName","organismID","collectionCode","catalogNumber") ]
Will return:
basisOfRecord scientificName occurrenceID collectionCode catalogNumber
1 PreservedSpecimens Duckeodendron cestroides 3456-A - Oliveira -INPA.12345A INPA 12345A
2 PreservedSpecimens Duckeodendron cestroides 3456-A - Oliveira -MO.ANOTHER FAKE CODE MO ANOTHER FAKE CODE
3 PreservedSpecimens Duckeodendron cestroides 3456-A - Oliveira -NY.574635 NY 574635
Import Vouchers for Existing Individuals
- Vouchers are samples of Individuals deposited in BioCollections. If the Biocollection is a genetic resource, the Voucher may just represent a DNA or tissue sample.
- Therefore, Vouchers must belong to an Individual and to a BioCollection, which are the mandatory information to register vouchers. Collectors, collection date, location and taxonomic identity may just be extracted from the Individual’s record.
- A Voucher may have its own collector, collector number and collecting date, different from the Individual it belongs to, when needed.
The mandatory fields are:
individual= individual id or fullname (organismID);biocollection= acronym or id of the BioCollection - useodb_get_biocollections()to check if it is registered, otherwise, first store the BioCollection in in the database;
For additional fields see Voucher POST API.
A simple voucher import
#a holotype voucher with same collector and date as individual
onevoucher = data.frame(individual=1,biocollection="INPA",biocollection_number=1234,biocollection_type=1,dataset=1)
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ" #this must be your token not this value
cfg = odb_config(base_url=base_url, token = token)
odb_import_vouchers(onevoucher, odb_cfg=cfg)
#get the imported voucher
voucher = odb_get_vouchers(params=list(individual=1),cfg)
vouchers[,c("basisOfRecord","scientificName","occurrenceID","collectionCode","catalogNumber") ]
Different voucher for an individual
Two vouchers for the same individual, one with the same collector and date as the individual, the other at different time and by other collectors.
#one with same date and collector as individual
one = data.frame(individual=2,biocollection="INPA",biocollection_number=1234,dataset=1,collector=NA,number=NA,date=NA)
#this one with different collector and date
two= data.frame(individual=2,biocollection="INPA",biocollection_number=4435,dataset=1,collector="Oliveira, A.A. de|João Batista da Silva",number=3456,date="1991-08-01")
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ" #this must be your token not this value
cfg = odb_config(base_url=base_url, token = token)
to.odb = rbind(one,two)
odb_import_vouchers(to.odb, odb_cfg=cfg)
#get the imported voucher
voucher = odb_get_vouchers(params=list(individual=2),cfg)
voucher[,c("basisOfRecord","scientificName","occurrenceID","collectionCode","catalogNumber") ]
Output of imported records:
basisOfRecord scientificName occurrenceID collectionCode catalogNumber recordedByMain
1 PreservedSpecimens Unidentified plot tree - Vicentini -INPA.1234 INPA 1234 Vicentini, A.
2 PreservedSpecimens Unidentified 3456 - Oliveira -INPA.4435 INPA 4435 Oliveira, A.A. de
6.2.9 - Import Measurements
- Measurement have one restriction: duplicated measurement values or categories for the same object_id and same date will not be imported unless you specify a
duplicated=1with for the record. - Measurements are straightforward, but you need to grap some info from other models to import them. You need to provide a
datasetfor which you do have permissions,trait_id(export_name or id),value(content varies depending on trait type),date(a complete or incomplete date),person(person that measured),object_type(one of taxon, individual, voucher or location) andobject_id(its odb id). For LINK trait type you must provide alink_idcolumn, and ‘value’ becomes optional for this specific case. - TIP: To link among measurements for different traits you may add standard keys to the
notemeasurement field. For example, you measure LeafLength and LeafWidth for the same leaves, so in both measurements add “leaf1” ,“leaf2” to the note field to link them. Also, if you measure DBH and DBH.POM in a forest plot inventory and have several stems for a plant, you may add “stem1”,“stem2” to permit matching a stem DBH to its specific height of measurement. This kind of link may be useful in analyses. - Several validations will happen during measurement importation. Consider verifying the data locally before submitting a job to facilitate troubleshooting, and guarantee the values are consistent with your expectations. You will not be able to UPDATE nor DELETE measurements with the API, so, avoid importing several measurements that will require fixing afterwards (one by one).
- For Quantitative Traits, OpenDataBio cannot validate that your measurements are in the same unit as the Trait definition. Therefore, check which unit the trait has, and convert your data accordingly before uploading.
Measurements can be imported using odb_import_measurements(). Read carefully the Measurements POST API.
Quantitative measurements
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ" #this must be your token not this value
cfg = odb_config(base_url=base_url, token = token)
#get the trait id from the server (check that trait exists)
#generate some fake data for 10 measurements
dbhs = sample(seq(10,100,by=0.1),10)
object_ids = sample(1:3,length(dbhs),replace=T)
dates = sample(as.Date("2000-01-01"):as.Date("2000-03-31"),length(dbhs))
dates = lapply(dates,as.Date,origin="1970-01-01")
dates = lapply(dates,as.character)
dates = unlist(dates)
to.odb = data.frame(
trait_id = 'dbh',
value = dbhs,
date = dates,
object_type = 'Individual',
object_id=object_ids,
person="Oliveira, A.A. de",
dataset = 1,
notes = "some fake measurements",
stringsAsFactors=F)
#this will only work if the person exists, the individual ids exist
#and if the trait with export_name=dbh exist
odb_import_measurements(to.odb,odb_cfg=cfg)
Get the imported data:
dad = odb_get_measurements(params = list(dataset=1),odb_cfg=cfg)
dad[,c("id","basisOfRecord", "measured_type", "measured_id", "measurementType",
"measurementValue", "measurementUnit", "measurementDeterminedDate",
"datasetName", "license")]
id basisOfRecord measured_type measured_id measurementType measurementValue measurementUnit measurementDeterminedDate
1 1 MeasurementsOrFact App\\Models\\Individual 3 dbh 86.8 centimeters 2000-02-19
2 2 MeasurementsOrFact App\\Models\\Individual 2 dbh 84.8 centimeters 2000-03-25
3 3 MeasurementsOrFact App\\Models\\Individual 2 dbh 65.7 centimeters 2000-03-15
4 4 MeasurementsOrFact App\\Models\\Individual 3 dbh 88.0 centimeters 2000-03-05
5 5 MeasurementsOrFact App\\Models\\Individual 3 dbh 35.3 centimeters 2000-01-04
6 6 MeasurementsOrFact App\\Models\\Individual 2 dbh 36.0 centimeters 2000-03-23
7 7 MeasurementsOrFact App\\Models\\Individual 2 dbh 78.6 centimeters 2000-03-22
8 8 MeasurementsOrFact App\\Models\\Individual 2 dbh 69.7 centimeters 2000-03-09
9 9 MeasurementsOrFact App\\Models\\Individual 3 dbh 12.3 centimeters 2000-01-30
10 10 MeasurementsOrFact App\\Models\\Individual 3 dbh 14.7 centimeters 2000-01-18
datasetName license
1 Dataset test CC-BY 4.0
2 Dataset test CC-BY 4.0
3 Dataset test CC-BY 4.0
4 Dataset test CC-BY 4.0
5 Dataset test CC-BY 4.0
6 Dataset test CC-BY 4.0
7 Dataset test CC-BY 4.0
8 Dataset test CC-BY 4.0
9 Dataset test CC-BY 4.0
10 Dataset test CC-BY 4.0
Categorical measurements
Categories MUST be informed by their ids or name in the value field. For CATEGORICAL or ORDINAL traits, value must be single value. For CATEGORICAL_MULTIPLE, value may be one or multiple categories ids or names separated by one of | or ; or ,.
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ" #this must be your token not this value
cfg = odb_config(base_url=base_url, token = token)
#a categorical trait
(odbtraits = odb_get_traits(params=list(name="specimenFertility"),odb_cfg = cfg))
#base line
to.odb = data.frame(trait_id = odbtraits$id, date = '2021-07-31', stringsAsFactors=F)
#the plant was collected with both flowers and fruits, so the value are the two categories
value = c("Flowers","Fruits")
#get categories for this trait if found
(cats = odbtraits$categories[[1]])
#check that your categories are registered for the trait and get their ids
value = cats[match(value,cats$name),'id']
#make multiple categories ids a string value
value = paste(value,collapse=",")
to.odb$value = value
#this links to a voucher
to.odb$object_type = "Voucher"
#get voucher id from API (must be ID).
#Search for collection number 1234
odbspecs = odb_get_vouchers(params=list(number="3456-A"),odb_cfg=cfg)
to.odb$object_id = odbspecs$id[1]
#get dataset id
odbdatasets = odb_get_datasets(params=list(name='Dataset test'),odb_cfg=cfg)
head(odbdatasets)
to.odb$dataset = odbdatasets$id
#person that measured
odbperson = odb_get_persons(params=list(search='ana cristina sega'),odb_cfg=cfg)
to.odb$person = odbperson$id
#import'
odb_import_measurements(to.odb,odb_cfg=cfg)
#get imported
dad = odb_get_measurements(params = list(voucher=odbspecs$id[1]),odb_cfg=cfg)
dad[,c("id","basisOfRecord", "measured_type", "measured_id", "measurementType",
"measurementValue", "measurementUnit", "measurementDeterminedDate",
"datasetName", "license")]
id basisOfRecord measured_type measured_id measurementType measurementValue measurementUnit
1 11 MeasurementsOrFact App\\Models\\Voucher 1 specimenFertility Flowers, Fruits NA
measurementDeterminedDate datasetName license
1 2021-07-31 Dataset test CC-BY 4.0
Color measurements
For color values you have to enter color as their hex RGB strings codes, so they can be rendered graphically and in the web interface. Therefore, any color value is allowed, and it would be easier to use the palette colors in the web interface to enter such measurements. Package gplots allows you to convert color names to hex RGB codes if you want to do it through the API.
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ" #this must be your token not this value
cfg = odb_config(base_url=base_url, token = token)
#get the trait id from the server (check that trait exists)
odbtraits = odb_get_traits(odb_cfg=cfg)
(m = match(c("fruitColor"),odbtraits$export_name))
#base line
to.odb = data.frame(trait_id = odbtraits$id[m], date = '2014-01-13', stringsAsFactors=F)
#get color value
#install.packages("gplots",dependencies = T)
library(gplots)
(value = col2hex("red"))
to.odb$value = value
#this links to a specimen
to.odb$object_type = "Individual"
#get voucher id from API (must be ID). Search for collection number 1234
odbind = odb_get_individuals(params=list(tag='3456'),odb_cfg=cfg)
odbind$scientificName
to.odb$object_id = odbind$id[1]
#get dataset id
odbdatasets = odb_get_datasets(params=list(name='Dataset test'),odb_cfg=cfg)
head(odbdatasets)
to.odb$dataset = odbdatasets$id
#person that measured
odbperson = odb_get_persons(params=list(search='ana cristina sega'),odb_cfg=cfg)
to.odb$person = odbperson$id
odb_import_measurements(to.odb,odb_cfg=cfg)

Database link type measurements
The LINK trait type allows one to register count data, as for example the number of individuals of a species at a particular location. You have to provide the linked object (link_id), which may be a Taxon or a Voucher depending on the trait definition, and then value recieves the numeric count.
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ" #this must be your token not this value
cfg = odb_config(base_url=base_url, token = token)
#get the trait id from the server (check that trait exists)
odbtraits = odb_get_traits(odb_cfg=cfg)
(m = match(c("taxonCount"),odbtraits$export_name))
#base line
to.odb = data.frame(trait_id = odbtraits$id[m], date = '2014-01-13', stringsAsFactors=F)
#the taxon to link the count value
odbtax = odb_get_taxons(params=list(name='Ocotea guianensis'),odb_cfg=cfg)
to.odb$link_id = odbtax$id
#now add the count value for this trait type
#this is optional for this measurement,
#however, it would make no sense to include such link without a count in this example
to.odb$value = 23
#a note to clarify the measurement (optional)
to.odb$notes = 'No voucher, field identification'
#this measurement will link to a location
to.odb$object_type = "Location"
#get location id from API (must be ID).
#lets add this to a transect
odblocs = odb_get_locations(params=list(adm_level=101,limit=1),odb_cfg=cfg)
to.odb$object_id = odblocs$id
#get dataset id
odbdatasets = odb_get_datasets(params=list(name='Dataset test'),odb_cfg=cfg)
head(odbdatasets)
to.odb$dataset = odbdatasets$id
#person that measured
odbperson = odb_get_persons(params=list(search='ana cristina sega'),odb_cfg=cfg)
to.odb$person = odbperson$id
odb_import_measurements(to.odb,odb_cfg=cfg)

Spectral measurements
value must be a string of spectrum values separated by “;”. The number of concatenated values must match the Trait value_length attribute of the trait, which is extracted from the wavenumber range specification for the trait. So, you may easily check this before importing with odb_get_traits(params=list(fields='all',type=9),cfg)
library(opendatabio)
base_url="https://opendb.inpa.gov.br/api"
token ="GZ1iXcmRvIFQ" #this must be your token not this value
cfg = odb_config(base_url=base_url, token = token)
#read a spectrum
spectrum = read.table("1_Sample_Planta-216736_TAG-924-1103-1_folha-1_abaxial_1.csv",sep=",")
#second column are NIR leaf absorbance values
#the spectrum has 1557 values
nrow(spectrum)
#[1] 1557
#collapse to single string
value = paste(spectrum[,2],collapse = ";")
substr(value,1,100)
#[1] "0.6768057;0.6763237;0.6755353;0.6746023;0.6733549;0.6718447;0.6701176;0.6682984;0.6662288;0.6636459;"
#get the trait id from the server (check that trait exists)
odbtraits = odb_get_traits(odb_cfg=cfg)
(m = match(c("driedLeafNirAbsorbance"),odbtraits$export_name))
#see the trait
odbtraits[m,c("export_name", "unit", "range_min", "range_max", "value_length")]
#export_name unit range_min range_max value_length
#6 driedLeafNirAbsorbance absorbance 3999.64 10001.03 1557
#must be true
odbtraits$value_length[m]==nrow(spectrum)
#[1] TRUE
#base line
to.odb = data.frame(trait_id = odbtraits$id[m], value=value, date = '2014-01-13', stringsAsFactors=F)
#this links to a voucher
to.odb$object_type = "Voucher"
#get voucher id from API (must be ID).
#search for a collection number
odbspecs = odb_get_vouchers(params=list(number="3456-A"),odb_cfg=cfg)
to.odb$object_id = odbspecs$id[1]
#get dataset id
odbdatasets = odb_get_datasets(params=list(name='Dataset test'),odb_cfg=cfg)
to.odb$dataset = odbdatasets$id
#person that measured
odbperson = odb_get_persons(params=list(search='adolpho ducke'),odb_cfg=cfg)
to.odb$person = odbperson$id
#import
odb_import_measurements(to.odb,odb_cfg=cfg)
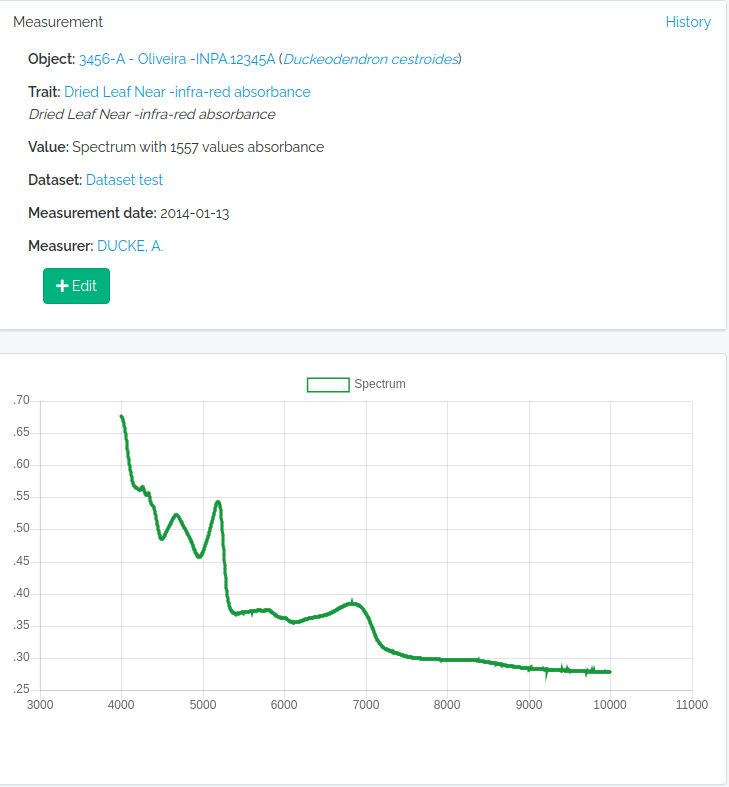
Text measurements
Just add the text to the value field and proceed as for the other trait types.